Bioutils: Difference between revisions
imported>Weigang |
imported>Weigang |
||
| Line 64: | Line 64: | ||
<div class="NavFrame"> | <div class="NavFrame"> | ||
<div class="NavHead">PerlDoc for '''bioseq'''</div> | <div class="NavHead">PerlDoc for '''bioseq'''</div> | ||
<div class="NavContent"> | <div class="NavContent" style="display: none;"> | ||
<h1 id="NAME">NAME</h1> | <h1 id="NAME">NAME</h1> | ||
<p>bioseq - Fasta sequence editing module based on BioPerl.</p> | <p>bioseq - Fasta sequence editing module based on BioPerl.</p> | ||
Revision as of 19:01, 29 September 2014
BioPerl-based Sequence Utilities
What is bioutils?
bioutils is a suite of Perl scripts that provide convenient command-line access to popular BioPerl methods. Designed as UNIX utilities, these tools aim to circumvent a constant need (and urge) for composing one-off BioPerl scripts for routine manipulations of sequences, alignments and trees.
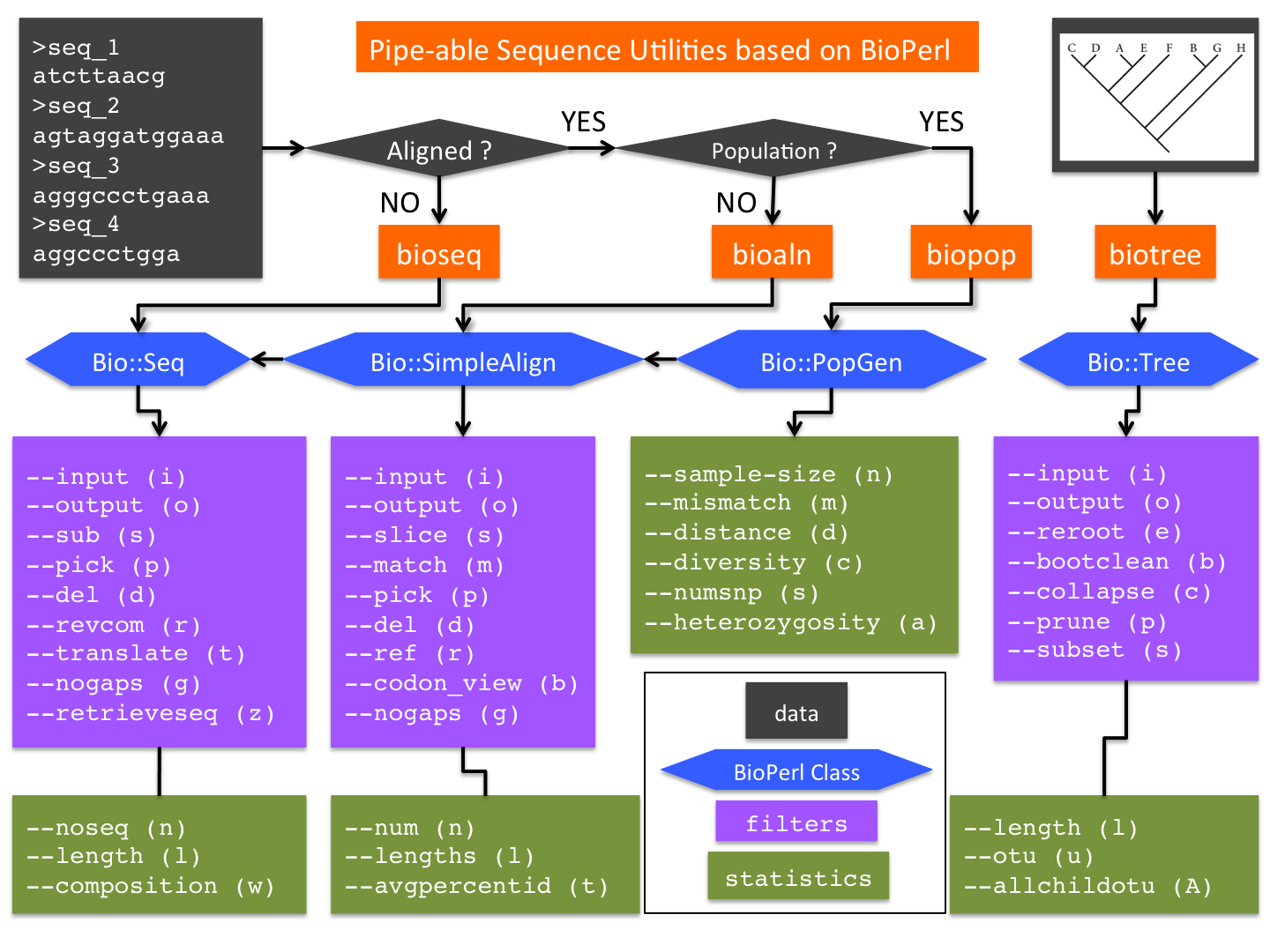
The initial release of bioutils consists of four utilities (Figure 1):
- bioseq: a wrapper of BioPerl class Bio::Seq (with additional methods)
- bioaln: a wrapper of Bio::SimpleAlign (which inherits Bio::Seq; with additional methods)
- biopop: a wrapper of Bio::PopGen (which can be converted from Bio::SimpleAlign; with additional methods)
- biotree: a wrapper of Bio::tree (with additional methods)
These utilities have been in development since 2002 in the lab of Dr Weigang Qiu at Hunter College of the City University of New York. They are the main code base of the Qiu Lab, which specializes in microbial evolutionary genomics. They proved to be convenient, efficient, and popular among students and researchers passing through the lab. By releasing bioutils as an Open Source tool (perhaps as a part of bioperl distribution), we hope to (1) share our experience and (2) invite other developers to join the effort of making BioPerl more accessible.
Live Demos
Basic Usage
- bioseq
bioseq -l foo.fasta # print seq names and lengths from FASTA (default format) file
bioseq -r foo.fasta # reverse complement
bioseq -t1 foo.fasta # translate in the +1 frame
bioseq -t3 foo.fasta # translate in +1, +2, and +3 frames
bioseq -t6 foo.fasta # translate in all 6 frames
bioseq -p'id:seq_1' foo.fasta # pick a sequence by ID
bioseq -p'order:3' # pick the 3rd sequence
bioseq -p're:Human' foo.fasta # pick all sequences labeled as "Human" (by regular expression)
bioseq -g foo.fasta # remove all gaps
bioseq -z'CP003201' -o'genbank' # retrieve a GenBank file with accession
bioseq -z'CP003201' -o'fasta' # same file in FASTA- bioaln
bioaln -i'fasta' -o'phylip' foo.fasta # convert a FASTA alignment to PHYLIP
bioaln -l foo.aln # print alignment length of a CLUSTALW (default format) file
bioaln -s'100, 200' foo.aln # obtain an alignment slice
bioaln -m foo.aln # show only variable sites
bioaln -r'seq_2' foo.aln # use "seq_2" as reference (first) sequence
bioaln -g foo.aln # remove gapped sites
bioaln -p'seq_1,seq_3,seq_6' foo.aln # pick a subset of sequences
bioaln -d'seq_1,seq_3,seq_6' foo.aln # delele a subset of sequences- biotree
- biopop
Power usage (with pipes)
# Pipe with the same utility
bioseq -p'order:5' foo.fasta | bioseq -s'100,200' | bioseq -r | bioseq -t1 # pick, subseq, revcom, and translate
# Pipe among utilities
bioaln -o'fasta' foo.aln | bioseq -g # remove gaps within individual sequencesCreative usage (with BASH utils)
echo -ne ">lookup\nATG\n" | bioseq -t1 # Lookup a codon product
len=$(bioaln -l foo.aln); len_degap=$(bioaln -g foo.aln | bioaln -l); echo "$len-$len_degap" | bc -l # count alignment gapsFull documentation
Release 1.0 Notes
- Installation
- Dependency
Main contributors
- Yozen Hernandez
- Levy Vargas
- Pedro Pagan
- Che Martin
- James Haven
- Girish Ramrattan
- Raymond Liang
- Saymon Akther
- Daniel Packer
- Weigang Qiu