EEB BootCamp 2020: Difference between revisions
Jump to navigation
Jump to search
Bioinformatics Boot Camp for Ecology & Evolution: Genomic Epidemiology
Thursday, Aug 6, 2020, 2 - 3:30pm
Instructors: Dr Weigang Qiu & Ms Saymon Akther
Email: weigang@genectr.hunter.cuny.edu
Lab Website: http://diverge.hunter.cuny.edu/labwiki/
imported>Lab No edit summary |
imported>Lab No edit summary |
||
| Line 92: | Line 92: | ||
bcftools view --exclude-types indels calls.bcf > snps.vcf | bcftools view --exclude-types indels calls.bcf > snps.vcf | ||
## filter sites by allele counts: only | ## filter sites by allele counts: only keep informative sites | ||
vcftools --vcf snps.vcf --mac 2 --recode --recode-INFO-all --out snps2.vcf | vcftools --vcf snps.vcf --mac 2 --recode --recode-INFO-all --out snps2.vcf | ||
Revision as of 04:48, 4 August 2020
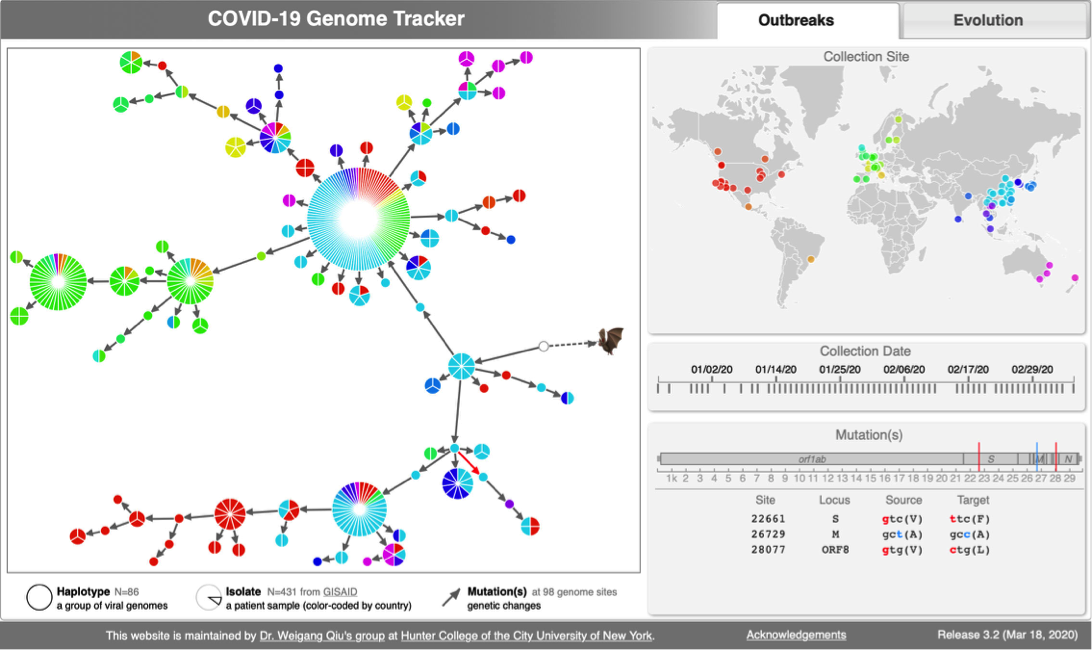
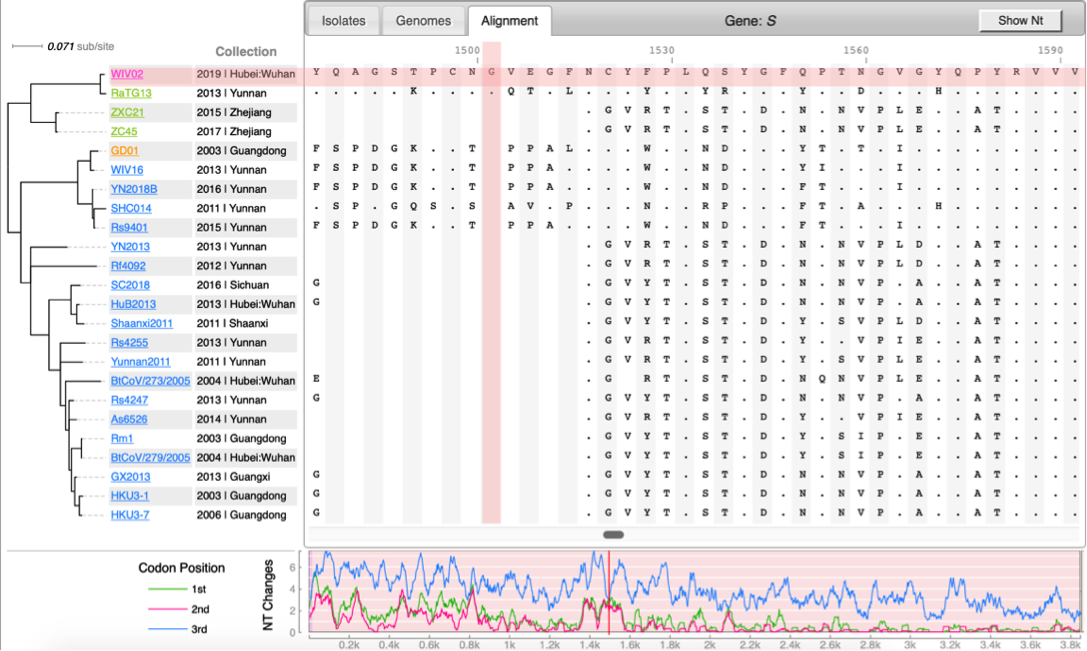
| CoV Genome Tracker | Coronavirus evolutuon | Lyme Disease (Borreliella) |
|---|---|---|
Case studies
- Is it necessary to mention nextstrain?
Bioinformatics tools for genomic epidemiology
Required for the tutorial
- bcftools: Reading/writing BCF2/VCF/gVCF files and calling/filtering/summarising SNP and short indel sequence variants Installation link
- vcftools: To work with genetic variation data in the form of VCF files Github link
- Sequence format converter Web tool
- TCS: To infer Haplotype network, TCS.jar file is provided, Required Java. PubMed link
- Web-interactive visualization of Haplotype Network with tcsBU Web tool; Paper
- ggplot (R)
Not required for the tutorial. Recommended
- BpWrapper: command-line tools for manipulation of sequences, alignment, and tree (based on BioPerl). Github Link; Flowchart from publication
- Pairwise genome alignment with MUMMER: Github link
- Samtools: Reading/writing/editing/indexing/viewing SAM/BAM/CRAM format Installation link
CoV genome data set
- N=100 SARS-CoV-2 genomes collected during January, February & March 2020. Data source & acknowledgement GIDAID (Warning: You need to acknowledge GISAID if you reuse the data in any publication)
- Download the folder "bootcamp_august_6th_2020": data file
- unzip the folder
unzip bootcamp_august_6th_2020.zip
- View files
ls -lrt # long list, in reverse timeline
ls cov_data # a folder of 100 CoV2 genomes in FASTA format, pairwise genome alignment sam and indexed sorted bam files generated by bwa (or nucmer) and samtools
# We skipped bwa (or nucmer) and samtools part of the tutorial for time constrain. The bash script used to generate these files is available on request
ls cov_data/*sorted.bam | wc # 100 sorted.bam files correspond to 100 sequence files
less ref.fas # NC_045512 as reference sequence, "q" to quit
less metadata_cov.txt # a tsv file that contains collection dates and geographic information of 100 CoV2 genomes
wc metadata_cov.txt
file TCS.jar # Java application
less bcf-snp-call.sh # a file contain all the bash commands required to call SNPs and generate vcf file of 100 CoV2 genomes
less ploidy.txt # to specify the ploidy=1 during vcf SNP call
less rgb.txt #rgb color code to color the phylogenetic network
- Additional files
Tutorial
- 2-2:30: Introduction on pathogen phylogenomics
- 2:30-2:55: Part 1: Calling SNPs and creating VCF file
## Create alignment pileup and call variants using plodity file(plodity 1), multiallelic, first output is bam then piped to bcf ##
bcftools mpileup -Ou -f ref.fas cov_data/*sorted.bam | bcftools call -mv --ploidy-file ploidy.txt -Ob -o calls.bcf
## Get Stats, check # of records (SNPs), # of indels, TS/TV ratio
##( expeced more transitions than transversions) #for cov2 should be around more or less 2.5
bcftools stats calls.bcf | less
#remove indels and save into vcf format
bcftools view --exclude-types indels calls.bcf > snps.vcf
## filter sites by allele counts: only keep informative sites
vcftools --vcf snps.vcf --mac 2 --recode --recode-INFO-all --out snps2.vcf
## rename the snp2 recode file
mv snps2.vcf.recode.vcf snps2.vcf
#Get stats, check # of records (informative SNPs) and TS/TV ratio (the ratio will increase)
bcftools stats snps2.vcf | less
## Rename the samples
bcftools query -l snps2.vcf > samples
wc samples
less samples
# make sure you don't have exsiting rename_ids.txt file since we are going to append on the file
if [ -e rename_ids.txt ];then rm rename_ids.txt ; fi
cat samples | while read line; do basename $line .sorted.bam | sed "s/Chromosome_//" >> rename_ids.txt ; done;
wc rename_ids.txt
#change the name of the samples
bcftools reheader -s rename_ids.txt snps2.vcf > snps3.vcf
less snps3.vcf
#create fasta file of snps
cat rename_ids.txt | while read line; do echo ">$line"; bcftools query -s "$line" -f '[%TGT]' snps3.vcf; echo; done > cov_haplotypes.fas
grep ">" cov_haplotypes.fas | wc
- 2:55-3:10: Part 2: Build and interactive visualize haplotype network with TCS and tcsBU
# Data pre-processing
# 1. Download genomes & meta data from GISAID
# 2. Run dnadist against a reference genome
man nucmer
dnadiff -h
dnadiff ref.fas <query FASTA>
mkdir fasta-files
cd fasta-files
for f in *.fas; do dnadiff ref.fas $f; done
<to be added: plot in R seq diff vs collection date>
# 3. Remove mis-assembled and reverse-complemented genomes
bioseq -d'file:'
# 4. Remove genomes with more than 10 non-ATCG bases
bioseq -d'ambig:10'
# 5. Run mafft (not run; takes too long)
# 6. Run snp-sites
snp-sites
java -jar -Xmx1g TCS.jar
- 3:10-3:25: Part 3: Visualization of SNPs frequency with vcftools and ggplot2
- Load graph file
- Load group file
- Load haplotype file
- 3:25-3:30: Q & A