BioMed-R-2020: Difference between revisions
imported>Weigang m (→March 7, 2019) |
imported>Weigang m (→March 14, 2019) |
||
| Line 157: | Line 157: | ||
===March 14, 2019=== | ===March 14, 2019=== | ||
* | * Mid-term exam (50 pts). Open Boook | ||
===March 22, 2019=== | ===March 22, 2019=== | ||
* R tutorial: Section 5.3. t-test | * R tutorial: Section 5.3. t-test | ||
Revision as of 19:32, 27 February 2020
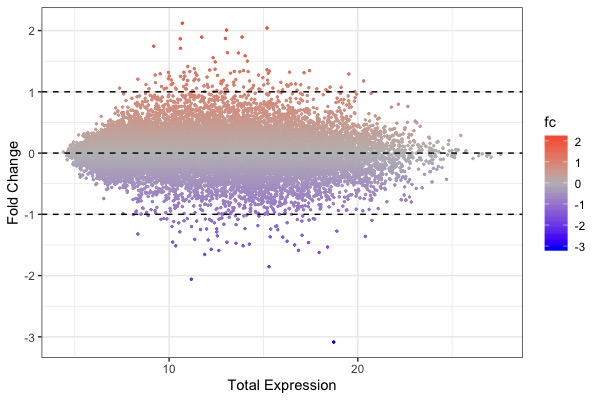
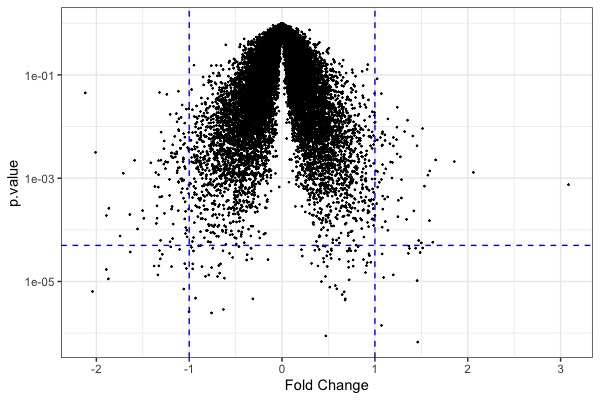
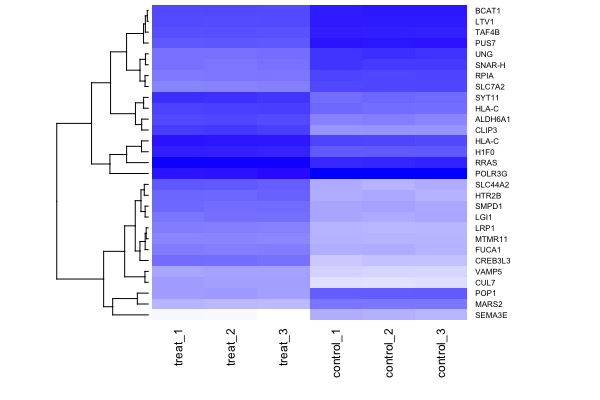
| MA plot | Volcano plot | Heat map |
|---|---|---|
Course Overview
Welcome to Introductory BioMedical Genomics, a seminar course for advanced undergraduates and graduate students. A genome is the total genetic content of an organism. Driven by breakthroughs such as the decoding of the first human genome and rapid DNA and RNA-sequencing technologies, biomedical sciences are undergoing a rapid & irreversible transformation into a highly data-intensive field, that requires familiarity with concepts in both biology, computational, and data sciences.
Genome information is revolutionizing virtually all aspects of life sciences including basic research, medicine, and agriculture. Meanwhile, use of genomic data requires life scientists to be familiar with concepts and skills in biology, computer science, as well as statistics.
This workshop is designed to introduce computational analysis of genomic data through hands-on computational exercises. Students are expected to be able to replicate key results of data analysis from published studies.
The pre-requisites of the course are college-level courses in molecular biology, cell biology, and genetics. Introductory courses in computer programming and statistics are preferred but not strictly required.
Learning goals
By the end of this course successful students will be able to:
- Describe next-generation sequencing (NGS) technologies & contrast it with traditional Sanger sequencing
- Explain applications of NGS technology including pathogen genomics, cancer genomics, human genomic variation, transcriptomics, meta-genomics, epi-genomics, and microbiome.
- Visualize and explore genomics data using R & RStudio
- Replicate key results using a raw data set produced by a primary research paper
Web Links
- Install R base: https://cloud.r-project.org
- Install R Studio (Desktop version): http://www.rstudio.com/download
- Textbook: Introduction to R for Biologists
- Download: R datasets
- A reference book: R for Data Science (Wickharm & Grolemund)
Quizzes and Exams
Student performance will be evaluated by attendance, weekly assignments, quizzes, and a final report:
- Attendance & In-class participation: 50 pts
- Assignments: 5 x 10 = 50 pts
- Quizzes: 2 x 25 pts = 50 pts
- Mid-term: 50 pts
- Final presentation & report: 100 pts
Total: 300 pts
Tips for Success
To maximize the your experience we strongly recommend the following strategies:
- Follow the directions for efficiently, finding high-impact papers, reading science research papers and preparing presentations.
- Read the papers, watch required videos and do the exercises regularly, long before you attend class.
- Attend all classes, as required. Late arrival results in loss of points.
- Keep up with online exercises. Don’t wait until the due date to start tasks.
- Take notes or annotate slides while attending the lectures.
- Listen actively and participate in class and in online discussions.
- Review and summarize material within 24 hrs after class.
- Observe the deadlines for submitting your work. Late submissions incur penalties.
- Put away cell phones, do not TM, email or play computer games in class.
Hunter/CUNY Policies
- Policy on Academic Integrity
Hunter College regards acts of academic dishonesty (e.g., plagiarism, cheating on homework, online exercises or examinations, obtaining unfair advantage, and falsification of records and official documents) as serious offenses against the values of intellectual honesty. The College is committed to enforcing the CUNY Policy on Academic Integrity, and we will pursue cases of academic dishonesty according to the Hunter College Academic Integrity Procedures. Students will be asked to read this statement before exams.
- ADA Policy
In compliance with the American Disability Act of 1990 (ADA) and with Section 504 of the Rehabilitation Act of 1973, Hunter College is committed to ensuring educational parity and accommodations for all students with documented disabilities and/or medical conditions. It is recommended that all students with documented disabilities (Emotional, Medical, Physical, and/or Learning) consult the Office of AccessABILITY, located in Room E1214B, to secure necessary academic accommodations. For further information and assistance, please call: (212) 772- 4857 or (212) 650-3230.
- Syllabus Policy
Except for changes that substantially affect implementation of the evaluation (grading) statement, this syllabus is a guide for the course and is subject to change with advance notice, announced in class or posted on Blackboard.
Course Schedule
Feb 1, 2020
- Introduction
- R Tutorial 1: Use interface, basic operations, load data. Slides:
| Assignment 1 (10 pts; Due next class 2/8, in hard copy) |
|---|
PropertyName,Density_250m,Density_500m,Density_1000m HighbridgePark,0.006561319,0.009462031,0.010578611 BronxRiverParkway,0.001318749,0.001978858,0.002652118 CrotonaPark,0.009412087,0.01164712,0.01202321 ClaremontPark,0.016391948,0.019972485,0.020350481 VanCortlandtPark,0.000550151,0.000979312,0.001372675
|
Feb 8, 2019
- Introduction to NGS:
- 1-slide presentations on Next-Generation Sequencing Technologies (Group I)
- R Tutorial, Part 2. Data manipulation with dplyr. Slides:
| Assignment 2 (10 pts; Due next class 2/15, in hard copy) |
|---|
|
Feb 15, 2019
- NGS presentations (Group II)
- R Tutorial. Part 3. Data visualization with ggplot2. Slides:
- No assignment (go over slides and 3 tutorial scripts to prepare for Quiz next week)
Feb 22, 2019
- Quiz 1 (Open Book)
- R Tutorial: Part 4. BioStat (chi-square & t-test) Lecture slides:
| Assignment 3 (10 pts). In-class workshop. Evaluation of papers according to the following rubrics (submit by email) |
|---|
|
Feb 29, 2019
- Student submissions
| Student & project type | Citation & PubMed link | Research question | Samples, sample size, controls | Omics tech & NGS platform | Computational tools | Data visualization | Statistical tests | Data description & links |
|---|---|---|---|---|---|---|---|---|
| Tahir; microbiome | Kostic, A. D., et al. (2012). Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome research, 22(2), 292–298. PubMed | How does the composition of tumorous colorectal carcinoma tissue microbiome differ from non-tumorous adjacent tissue? | Colorectal carcinoma (Tumor) tumor tissue and non-tumorous adjacent nonneoplastic (Normal) tissue); 95 tumor/normal paired samples (190 total samples); Non tumorous adjacent nonneoplastic tissue as controls | 16S rDNA amplicon sequencing; 454 GS FLX Sequencing | Mothur | Bar plots, Boxplots, Scatterplots, Cladogram | Linear Discriminate Analysis (LDA) and Wilcox Rank Sum Test (non-parametric t-test) | NCBI Sequence Read Archive accession no. SRP000383. Pre-processed dataset can also be retrieved from R package, phyloseq: filepath = system.file("extdata", "study_1457_split_library_seqs_and_mapping.zip", package = "phyloseq"); kostic = microbio_me_qiime(filepath). The Kostic dataset is a phyloseq object (S4) consisting of sam_table, otu_table, table, phy_tree, and tax_table. Sample table includes metadata of samples collected including: Diagnosis, Race, Gender, etc. |
| Junho; Transcriptome | Gierlin ́ski M, Cole C, Schofield P, Schurch NJ, Sherstnev A, Singh V, Wrobel N, Gharbi K, Simpson G, Owen- Hughes T, Blaxter M, and Barton GJ. Statistical models for RNA-seq data derived from a two-condition 48-replicate experiment. Bioinformatics, 31(22):1–15, 2015. | These estimates are typically based on statistical models such as the negative binomial distribution, which is employed by the tools, edgeR, DESeq and cuffdiff. Until now, the validity of these models has usually been tested on either low-replicate RNA-seq data or simulations. | RNA-seq dataset to date that contains mRNA from 48 replicates of two S. cerevisiae populations: wildtype vs snf2 knock-out mutants | Illumina HiSeq 2000 | RStudio | scatterplot, boxplot, heatmap | t-test, Wald test (2 factors); LRT for multiple factor | ftp.sra.ebi.ac.uk/vol1/fastq/ERR458/ERR458493/ERR458493.fastq.gz; ERR458493.fastq.gz; WT_1_Aligned.sortedByCoord.out; WT_2_Aligned.sortedByCoord.out; SNF2_1_Aligned.sortedByCoord.out.bam.bai; SNF2_2_Aligned.sortedByCoord.out.bam.bai |
| Example | Example | Example | Example | Example | Example | Example | Example | Example |
- Paper evaluation & selection
- R Tutorial: Part 4. BioStat (regression & ANOVA)
March 7, 2019
- Self study & prepare for mid-term (no class)
March 14, 2019
- Mid-term exam (50 pts). Open Boook
March 22, 2019
- R tutorial: Section 5.3. t-test
- Group presentations (Data visualization)
March 28, 2019
- (Self study; No live class)
- Abstract (200 words; individualized; due 3/30)
- Review contingency test & two-sample t-test
- Generate preliminary graphs
March 30, 2019
- 20 pts Quiz on contingency test & two-sample t-test
- Group presentations (Show preliminary graphs)
- Material & Methods (due 4/6)
April 4, 2019
- 20 pts Quiz
- R tutorial: Section 5.4. Regression analysis
- Results (due 4/13)
- Tables to show the dataset you work on (not all, but a sample)
- Figures with legend (R methods, x & y-axis, conclusion)
- 1-paragraph summary of your results
April 18, 2019
- 20 pts Quiz. Regression analysis
- Background & Introduction (due 5/4)
April 25, 2019
- Final presentation I. Graded on:
- Objective (original & your own)
- Material & methods (original & your own)
- Results (your own)
- Conclusion (your own)
- Conclusion (due 5/11)
May 2, 2019
- Self study: Prepare your 10-slide presentation
- No class (instructor travels)
May 16, 2019, 9-1pm
- Final presentation
- May 22, 2018 (Wed, 5pm) Final Report Due (hard copy; n my office or in mailbox)