Summer 2021: Difference between revisions
Jump to navigation
Jump to search
imported>Weigang |
imported>Weigang |
||
| Line 20: | Line 20: | ||
* Reading list | * Reading list | ||
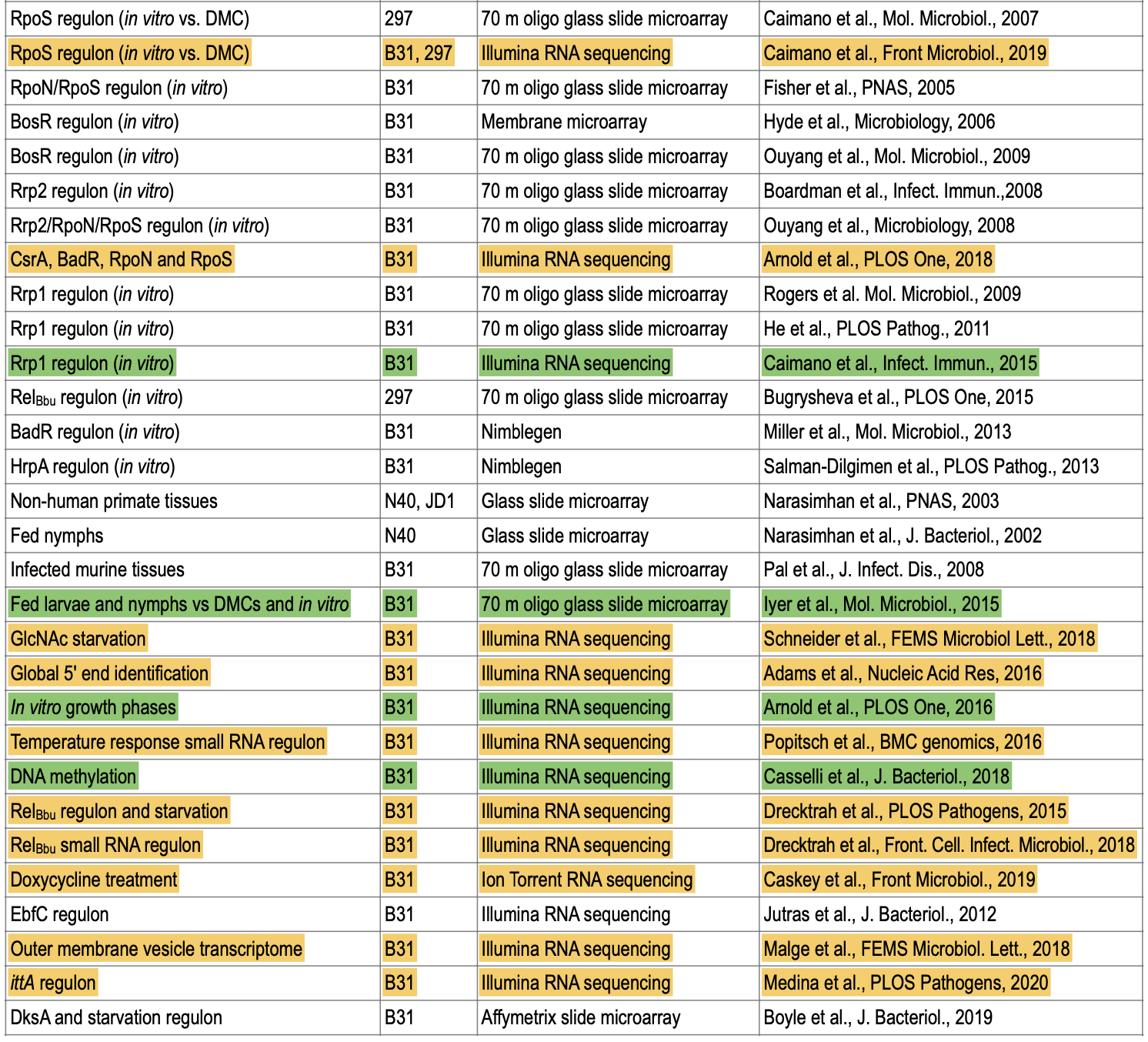
** Latest review book [https://www.caister.com/lyme Lyme Disease and Relapsing Fever Spirochetes: Genomics, Molecular Biology, Host Interactions and Disease Pathogenesis]. [https://www.caister.com/openaccess/pdf/9781913652616-05.pdf The chapter on gene regulation and transcriptomics] (notice Fig 1, Fig 2, and Table 1) | ** Latest review book [https://www.caister.com/lyme Lyme Disease and Relapsing Fever Spirochetes: Genomics, Molecular Biology, Host Interactions and Disease Pathogenesis]. [https://www.caister.com/openaccess/pdf/9781913652616-05.pdf The chapter on gene regulation and transcriptomics] (notice Fig 1, Fig 2, and Table 1) | ||
[[File:Transcriptome-table.png|thumbnail]] | |||
** Schward et al (2021). [https://pubmed.ncbi.nlm.nih.gov/33328355/ Multipartite Genome of Lyme Disease Borrelia: Structure, Variation and Prophages ] | ** Schward et al (2021). [https://pubmed.ncbi.nlm.nih.gov/33328355/ Multipartite Genome of Lyme Disease Borrelia: Structure, Variation and Prophages ] | ||
** Stevenson & Seshu (2018). [https://pubmed.ncbi.nlm.nih.gov/29064060/ Regulation of Gene and Protein Expression in the Lyme Disease Spirochete ] | ** Stevenson & Seshu (2018). [https://pubmed.ncbi.nlm.nih.gov/29064060/ Regulation of Gene and Protein Expression in the Lyme Disease Spirochete ] | ||
Revision as of 16:06, 15 June 2021
Group meeting/trip schedule
- June 3, 2021 (Thursday). Summer research kickoff
- June 8, 2021 (Tuesday). 11-2
- Algorithm development (Brian)
- NLP models of protein structure (Eamen, Roman, Edgar)
- Bb transcriptomics (Niemah & Jackie)
- HIV compartmentalization (Lily)
- June 10, 2021 (Thursday). No meeting. Field day
- Thursday: Desiree, Lily, Lia, Weigang
- Friday: John, Weigang
- Saturday: Weigang
- June 15, 2021 (Tuesday). 11-2
Project 1. Borrelia genomics
- Participants: Niemah, Jackie
- Questions & Goals:
- Upgrade database, genome pipeline, and website (Lia)
- Phylogeography & evolutionary maintenance of divided genome (Saymon)
- vls evolution (with simulation) & development of immunoflorescence microsopy methods(Lily). Live imaging.
- Reading list
- Latest review book Lyme Disease and Relapsing Fever Spirochetes: Genomics, Molecular Biology, Host Interactions and Disease Pathogenesis. The chapter on gene regulation and transcriptomics (notice Fig 1, Fig 2, and Table 1)
- Schward et al (2021). Multipartite Genome of Lyme Disease Borrelia: Structure, Variation and Prophages
- Stevenson & Seshu (2018). Regulation of Gene and Protein Expression in the Lyme Disease Spirochete
Project 2. Design algorithms for vaccines
- Participants: Dr Saad Mneimneih (CS Department), Brian
- Questions & Goals:
- Generalized algorithms for antigen with arbitrary tree shape
- Data set 1. Neutral evolution (with exponentially distributed branch lengths). Binary strings (L=100 bits) evolved from a coalescent tree of 20 leaves. Simulated with
rcoal(20); rTraitDisc; simSeq(). code from previous work - Data set 2. Two major clades. HA sequences from fluB
- Data set 3. Four major clades. Dengue
- Data set 4. Star-shaped tree, driven by recombination. OspC
- Data set 5. Multiple major clades. vls cassette in Lyme species
- Data set 1. Neutral evolution (with exponentially distributed branch lengths). Binary strings (L=100 bits) evolved from a coalescent tree of 20 leaves. Simulated with
- Combination algorithms
- Naive Bayes models to integrate immunogenicity data
- Natural language models to improve structural stability (see Project 4 below)
- Generalized algorithms for antigen with arbitrary tree shape
- Reading list
Project 3. HIV compartmentalized evolution
- Participants: Lily
- Questions and goals
- Do HIV evolve cell type tropisms within the host? Specifically, the Neural(N)-tropism vs T-cell(T)-tropism?
- Build a classifier of N-tropism HIV subtypes
- A presentation for an HIV conference in October
- Reading list
- HIV compartmentalized evolution: Evering et al (2014)
- Data sets
- ~500 sequences of env genes from 15 patients
- 2nd time point single-cell genome sequences for some of the patients
- Experimentally verified N-tropism subtypes
- Approach
- Evolutionary mechanisms: mutation, recombination, and adaptive selection
- Homoplasy index as a measure of compartmentalization? Randomization to obtain p-values of HI.
- Evolutionary rates & signature (BEAST)
- Tests of natural selection (PAML site models, branch-site models & MK analysis)
- Phylogenetic analysis: tree per individual; supertree; haplotype networks (per individual)
- Simulated compartmentalization
Project 4. Natural Language models of proteins
- Participants: Eamen, Roman, and Edgar
- Questions & Goals
- Learn, implement, and compare the existing tools
- Fine-tuning for OspC, to be integrated with the centroid algorithm
- 2nd-generation centroid design: k-means algorithm (with applications to vls, Dengue, flu B)
- Tools & Reading list
- Facebook ESM: pre-trained language models (for feature extraction)
- Step 1. implementation with colab
- Step 2. Fine-tuning with OspC seqs; extract embedding
- Step 3. Applications: classify (OspC vs VlsE), contact map (native vs synthetics), solubility
- Strodthoff et al (2020). Bioinformatics. UDSMProt: universal deep sequence models for protein classification. Source code on Github
- Rives et al (2021). PNAS. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Github repository
- Transformer: Vaswani et al (2017). Attention is All You Need Github repository for Huggingface/Transformer
- Facebook ESM: pre-trained language models (for feature extraction)