Monte Carlo Club: Difference between revisions
imported>Weigang |
imported>Lab |
||
| Line 31: | Line 31: | ||
## Which population can maintain genetic polymorphism (with two alleles) longer? | ## Which population can maintain genetic polymorphism (with two alleles) longer? | ||
## Which population gets fixed (only one allele remains) quicker? | ## Which population gets fixed (only one allele remains) quicker? | ||
{| class="wikitable sortable mw-collapsible" | |||
|- style="background-color:lightsteelblue;" | |||
! scope="col" style="width: 300px;" | Submitted Codes | |||
|- style="background-color:powderblue;" | |||
| | |||
*By John | |||
#Python | |||
<syntaxhighlight lang="python"> | |||
import numpy as np | |||
import sys | |||
from random import sample | |||
import matplotlib.pyplot as plt | |||
def simulator(gametes_rate, next_individuals, generations): | |||
individuals = [1 for i in range(50)] + [0 for j in range(50)] | |||
frequency = [] | |||
for generation in range(generations): | |||
gametes = [] | |||
for individual in individuals: | |||
gametes += [individual for i in range(gametes_rate)] | |||
individuals = sample(gametes, next_individuals) | |||
frequency.append(np.count_nonzero(individuals) / len(individuals)) | |||
return(frequency) | |||
N_100 = simulator(100, 100, 1000) | |||
N_1000 = simulator(100, 1000, 1000) | |||
# Create plots with pre-defined labels. | |||
# Alternatively,pass labels explicitly when calling `legend`. | |||
fig, ax = plt.subplots() | |||
ax.plot(N_100, 'r', label='N=100') | |||
ax.plot(N_1000, 'b', label='N=1000') | |||
# Add x, y labels and title | |||
plt.ylim(-0.1, 1.1) | |||
plt.xlabel("Generation") | |||
plt.ylabel("Frequency") | |||
plt.title("Wright-Fisher Model") | |||
# Now add the legend with some customizations. | |||
legend = ax.legend(loc='upper right', shadow=True) | |||
# The frame is matplotlib.patches.Rectangle instance surrounding the legend. | |||
frame = legend.get_frame() | |||
frame.set_facecolor('0.90') | |||
# Set the fontsize | |||
for label in legend.get_texts(): | |||
label.set_fontsize('large') | |||
for label in legend.get_lines(): | |||
label.set_linewidth(1.5) # the legend line width | |||
plt.show() | |||
#Scala | |||
import scala.util.Random | |||
import scala.collection.mutable.ListBuffer | |||
val gamete_rate = 100 | |||
val offspr_rate = 100 | |||
val generations = 1000 | |||
var frequency: List[Double] = List() | |||
var individuals = List.fill(50)(0) ++ List.fill(50)(1) | |||
for(generation <- 1 to generations){ | |||
var gametes = individuals.map(x => List.fill(gamete_rate)(x)).flatten | |||
individuals = Random.shuffle(gametes).take(offspr_rate) | |||
frequency = frequency :+ individuals.count(_ == 1).toDouble / offspr_rate | |||
} | |||
print(frequency) | |||
#Spark | |||
val gamete_rate = 100 | |||
val offspr_rate = 100 | |||
val generations = 300 | |||
var frequency: List[Double] = List() | |||
var individuals = sc.parallelize(List.fill(50)("0") ++ List.fill(50)("1")) | |||
for(generation <- 1 to generations){ | |||
val gametes = individuals.flatMap(x => (x * gamete_rate).split("").tail) | |||
individuals = sc.parallelize(gametes.takeSample(false, offspr_rate)) | |||
val count = individuals.countByValue | |||
frequency = frequency :+ count("1").toDouble / offspr_rate | |||
} | |||
print(frequency) | |||
</syntaxhighlight> | |||
*By Brian | |||
<syntaxhighlight lang="bash"> | |||
Wright<-function(pop.size,gam.size,generation) { | |||
if( pop.size%%2==0) { | |||
pop<-c(rep("A",pop.size*.5),rep("a",pop.size)) | |||
frequency<-.5 | |||
time<-0 | |||
geneticDrift<-data.frame(frequency,time) | |||
for (i in 1:generation) { | |||
largePop<-rep(pop,gam.size) | |||
samplePop<-sample(largePop,100) | |||
cases<-table(samplePop) | |||
if (cases[1]<100 && cases[2]<100 ) | |||
{propA<-(cases[1]/100) | |||
geneticDrift<-rbind(geneticDrift,c(propA,i)) | |||
pop<-c(rep("A",cases[2]),rep("a",cases[1])) | |||
} | |||
} | |||
plot(geneticDrift$frequency~geneticDrift$time,type="b", main="Genetic Drift N=1000", xlab="time",ylab="Proportion Pop A",col="red", pch=18 ) | |||
} | |||
if(pop.size%%2==1 ) { | |||
print("Initial Population should be even. Try again") | |||
} | |||
} | |||
Wright(1000,100,1000) | |||
</syntaxhighlight> | |||
*By Jamila | |||
<syntaxhighlight lang="bash"> | |||
Genetic_code = function(t,R){ | |||
N<- 100 | |||
p<- 0.5 | |||
frequency<-as.numeric(); | |||
for (i in 1:t){ | |||
A1=rbiom(1,2*N,p) | |||
p=A1/(N*2); | |||
frequency[length(frequency)+1]<-p; | |||
} | |||
plot(frequency, type="1",ylim=c(0,1),col=3,xlab="t",ylab=expression(p(A[1]))) | |||
} | |||
</syntaxhighlight> | |||
*By Sharon | |||
<syntaxhighlight lang="bash"> | |||
p=0.5 | |||
N=100 | |||
g=0 | |||
pop <- c(rep("A", 50), rep("a", 50)) #create a population of 2 alleles | |||
data<-data.frame(g,p) #a set of variables of the same # of rows | |||
for (i in 1:1000) { #generation | |||
gam_pl <-rep(pop, 100) #gamete pool | |||
gam_sam <-sample(gam_pl, 100) #sample from gamete pool | |||
tab_all<-table(gam_sam) #table ps sample | |||
all_freq<-tab_all[1]/100 #get allele frequency | |||
data<-rbind(data, c(i,all_freq)) | |||
pop<-c(rep("A",tab_all[1])) | |||
} | |||
plot(data$g,data$p) | |||
</syntaxhighlight> | |||
*By Sipa | |||
<syntaxhighlight lang="bash"> | |||
pop <- c(rep("A", 50), rep("a", 50)) | |||
wright_fisher <- function(N,gen_time,gam) { | |||
N=N | |||
gen_time = gen_time | |||
x = numeric(gen_time) | |||
x[1] = gam | |||
for (i in 2:1000) { | |||
k=(x[i-1])/N | |||
n=seq(0,N,1) | |||
prob=dbinom(n,N,k) | |||
x[i]=sample(0:N, 1, prob=prob) | |||
} | |||
plot(x[1:gen_time], type="l", pch=10) | |||
} | |||
pool <- rep(pop, times = 100) | |||
s_pool <- sample(pool, size = 100, replace = F) | |||
table(s_pool) | |||
wright_fisher2 <- function(pop_size,al_freq,gen_time) { | |||
pop <- c(rep("A", pop_size/2), rep("a", pop_size/2)) | |||
pool <-rep(pop, times=100) | |||
s_pool <- sample(pool, size = 100, replace = T) | |||
for(i in 1:gen_time) | |||
s_pool <- sample(sample(pool, size = pop_size, replace = F)) | |||
a.f <- table(s_pool)[1]/100 | |||
return(a.f) | |||
} | |||
</syntaxhighlight> | |||
*By Nicolette | |||
<syntaxhighlight lang="bash"> | |||
N=100 | |||
p=0.5 | |||
g=1000 | |||
sim=100 | |||
Genetic_drift=array(0, dim=c(g,sim)) | |||
Genetic_drift[1,]=rep(N*p,sim) | |||
for(i in 1:sim) { | |||
for(j in 2:g){ | |||
X[j,i]=rbinom(1,N,prob=X[j-1,i]/N) | |||
} | |||
} | |||
Genetic_drift=data.frame(X/N) | |||
matplot(1:1000, (X/N), type="l",ylab="allele_frequency",xlab="generations") | |||
</syntaxhighlight> | |||
== March 11, 2017 "Stoplights" (Due 3/18/2017)== | == March 11, 2017 "Stoplights" (Due 3/18/2017)== | ||
Revision as of 22:25, 2 April 2017
(This Week) March 31, 2017 "Drift-Mutation Balance" (Due 4/7/2017)
We will explore genetic drift of a DNA fragment with mutation under Wright-Fisher model. From last week's exercise, we conclude that a population will sooner or later lose genetic diversity, if no new alleles are generated (by e.g., mutation or migration).
Mutation, in contrast, increases genetic diversity over time. Under neutrality (no natural selection against or for any mutation, e.g., on an intron sequence), the population will reach an equilibrium point when the loss of genetic diversity by drift is cancelled out by increase of genetic diversity by mutation.
You job is to find this equilibrium point by simulation, given a population size (N) and a mutation rate (mu). The expected answer is pi=2N*mu, where pi is a measure of genetic diversity using DNA sequences, which is the average pairwise sequence differences within a population.
A suggested algorithm is:
- Start with a homogeneous population with N=100 identical DNA sequences (e.g., with a length of L=1e4 bases, or about 5 genes) [R hint:
dna <- sample(c("a", "t", "c", "g"), size=1e5, replace=T, prob = rep(0.25,4))] - Write a mutation function, which will mutate the DNA based on Poisson process (since mutation is a rare event). For example, if mu=1e-4 per generation per base per individual (too high for real, but faster for results to converge), then each generation the expected number of mutations would be L * mu = 1 per individual per generation for our DNA segment. You would then simulate the random number of mutations by using the R function
rpois(lambda=1). - Apply the mutation function for each individual DNA copy (a total of N=100) during gamete production (100 gametes for each individual) at each generation (for a total of G=1000 generations).
- For each generation, instead of counting allele frequencies (as last week's problem), you would need another function to calculate & output average pairwise differences among the 100 individuals.
- Finally, you would graph pi over generation.
March 18, 2017 "Genetic Drift" (Due 3/31/2017)
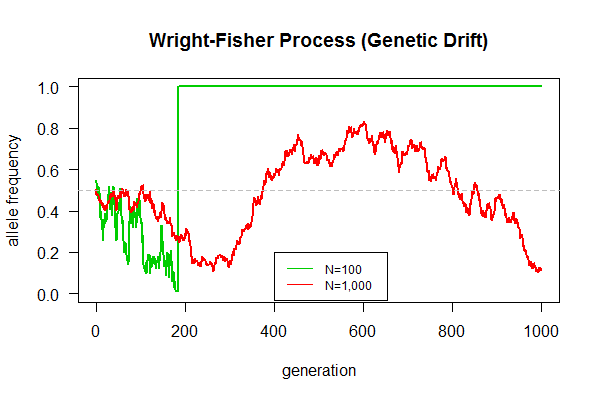
This is our first biological simulation. Mandatory assignment for all lab members (from interns to doctoral students). An expected result is shown in the graph.
Task: Simulate the Wright-Fisher model of genetic drift as follows:
- Begin with an allele frequency p=0.5, and a pop of N=100 haploid individuals [Hint:
pop <- c(rep("A", 50), rep("a", 50))] - Each individual produces 100 gametes, giving a total of 10,000 gametes [Hint: use a for loop with rep() function]
- Sample from the gamete pool another 100 to give rise to a new generation of individuals [Hint: use sample() function]
- Calculate allele frequency [Hint: use table() function]
- Repeat the above in succession for a total generation of g=1000 generations [Hint: create a function with three arguments, e.g., wright.fisher(pop.size, gamete.size, generation.time)]
- Plot allele frequency changes over generation time
- Be prepared to answer these questions:
- Why allele frequency fluctuate even without natural selection?
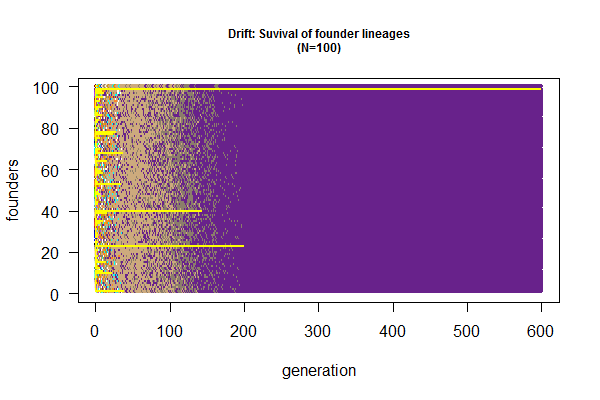
- What's the final fate of population, one allele left, or two alleles coexist indefinitely?
- Which population can maintain genetic polymorphism (with two alleles) longer?
- Which population gets fixed (only one allele remains) quicker?
| Submitted Codes | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
import numpy as np
import sys
from random import sample
import matplotlib.pyplot as plt
def simulator(gametes_rate, next_individuals, generations):
individuals = [1 for i in range(50)] + [0 for j in range(50)]
frequency = []
for generation in range(generations):
gametes = []
for individual in individuals:
gametes += [individual for i in range(gametes_rate)]
individuals = sample(gametes, next_individuals)
frequency.append(np.count_nonzero(individuals) / len(individuals))
return(frequency)
N_100 = simulator(100, 100, 1000)
N_1000 = simulator(100, 1000, 1000)
# Create plots with pre-defined labels.
# Alternatively,pass labels explicitly when calling `legend`.
fig, ax = plt.subplots()
ax.plot(N_100, 'r', label='N=100')
ax.plot(N_1000, 'b', label='N=1000')
# Add x, y labels and title
plt.ylim(-0.1, 1.1)
plt.xlabel("Generation")
plt.ylabel("Frequency")
plt.title("Wright-Fisher Model")
# Now add the legend with some customizations.
legend = ax.legend(loc='upper right', shadow=True)
# The frame is matplotlib.patches.Rectangle instance surrounding the legend.
frame = legend.get_frame()
frame.set_facecolor('0.90')
# Set the fontsize
for label in legend.get_texts():
label.set_fontsize('large')
for label in legend.get_lines():
label.set_linewidth(1.5) # the legend line width
plt.show()
#Scala
import scala.util.Random
import scala.collection.mutable.ListBuffer
val gamete_rate = 100
val offspr_rate = 100
val generations = 1000
var frequency: List[Double] = List()
var individuals = List.fill(50)(0) ++ List.fill(50)(1)
for(generation <- 1 to generations){
var gametes = individuals.map(x => List.fill(gamete_rate)(x)).flatten
individuals = Random.shuffle(gametes).take(offspr_rate)
frequency = frequency :+ individuals.count(_ == 1).toDouble / offspr_rate
}
print(frequency)
#Spark
val gamete_rate = 100
val offspr_rate = 100
val generations = 300
var frequency: List[Double] = List()
var individuals = sc.parallelize(List.fill(50)("0") ++ List.fill(50)("1"))
for(generation <- 1 to generations){
val gametes = individuals.flatMap(x => (x * gamete_rate).split("").tail)
individuals = sc.parallelize(gametes.takeSample(false, offspr_rate))
val count = individuals.countByValue
frequency = frequency :+ count("1").toDouble / offspr_rate
}
print(frequency)
Wright<-function(pop.size,gam.size,generation) {
if( pop.size%%2==0) {
pop<-c(rep("A",pop.size*.5),rep("a",pop.size))
frequency<-.5
time<-0
geneticDrift<-data.frame(frequency,time)
for (i in 1:generation) {
largePop<-rep(pop,gam.size)
samplePop<-sample(largePop,100)
cases<-table(samplePop)
if (cases[1]<100 && cases[2]<100 )
{propA<-(cases[1]/100)
geneticDrift<-rbind(geneticDrift,c(propA,i))
pop<-c(rep("A",cases[2]),rep("a",cases[1]))
}
}
plot(geneticDrift$frequency~geneticDrift$time,type="b", main="Genetic Drift N=1000", xlab="time",ylab="Proportion Pop A",col="red", pch=18 )
}
if(pop.size%%2==1 ) {
print("Initial Population should be even. Try again")
}
}
Wright(1000,100,1000)
Genetic_code = function(t,R){
N<- 100
p<- 0.5
frequency<-as.numeric();
for (i in 1:t){
A1=rbiom(1,2*N,p)
p=A1/(N*2);
frequency[length(frequency)+1]<-p;
}
plot(frequency, type="1",ylim=c(0,1),col=3,xlab="t",ylab=expression(p(A[1])))
}
p=0.5
N=100
g=0
pop <- c(rep("A", 50), rep("a", 50)) #create a population of 2 alleles
data<-data.frame(g,p) #a set of variables of the same # of rows
for (i in 1:1000) { #generation
gam_pl <-rep(pop, 100) #gamete pool
gam_sam <-sample(gam_pl, 100) #sample from gamete pool
tab_all<-table(gam_sam) #table ps sample
all_freq<-tab_all[1]/100 #get allele frequency
data<-rbind(data, c(i,all_freq))
pop<-c(rep("A",tab_all[1]))
}
plot(data$g,data$p)
pop <- c(rep("A", 50), rep("a", 50))
wright_fisher <- function(N,gen_time,gam) {
N=N
gen_time = gen_time
x = numeric(gen_time)
x[1] = gam
for (i in 2:1000) {
k=(x[i-1])/N
n=seq(0,N,1)
prob=dbinom(n,N,k)
x[i]=sample(0:N, 1, prob=prob)
}
plot(x[1:gen_time], type="l", pch=10)
}
pool <- rep(pop, times = 100)
s_pool <- sample(pool, size = 100, replace = F)
table(s_pool)
wright_fisher2 <- function(pop_size,al_freq,gen_time) {
pop <- c(rep("A", pop_size/2), rep("a", pop_size/2))
pool <-rep(pop, times=100)
s_pool <- sample(pool, size = 100, replace = T)
for(i in 1:gen_time)
s_pool <- sample(sample(pool, size = pop_size, replace = F))
a.f <- table(s_pool)[1]/100
return(a.f)
}
N=100
p=0.5
g=1000
sim=100
Genetic_drift=array(0, dim=c(g,sim))
Genetic_drift[1,]=rep(N*p,sim)
for(i in 1:sim) {
for(j in 2:g){
X[j,i]=rbinom(1,N,prob=X[j-1,i]/N)
}
}
Genetic_drift=data.frame(X/N)
matplot(1:1000, (X/N), type="l",ylab="allele_frequency",xlab="generations")
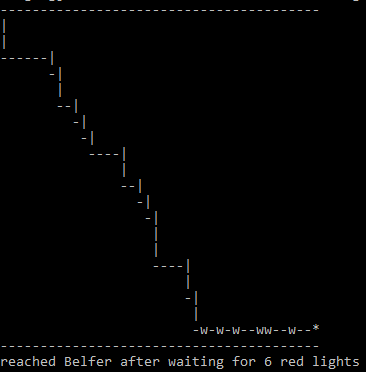
March 11, 2017 "Stoplights" (Due 3/18/2017)
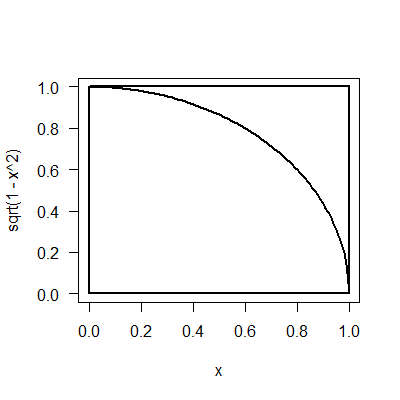
March 3, 2017 "PiE" (Due 3/10/2017)
Feb 24, 2017 "Idiots" (Due 3/3/2017)
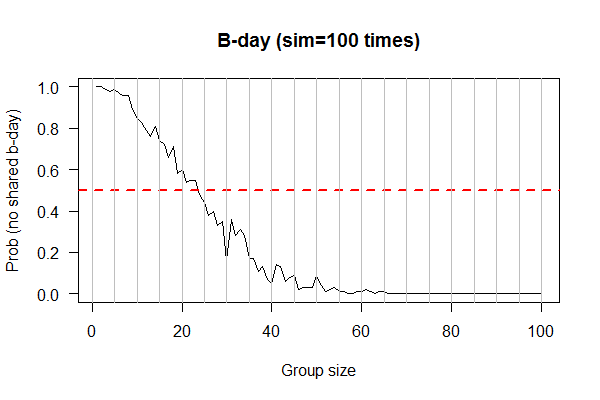
Feb 17, 2017 "Birthday" (Due 2/24/2017)
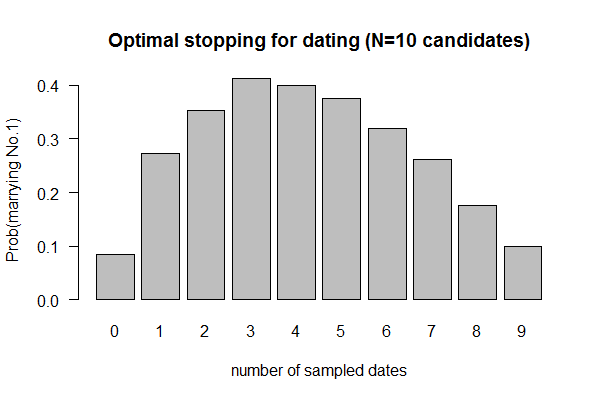
Feb 10, 2017 "Dating" (Valentine's Day Special; Due 2/17/2017)
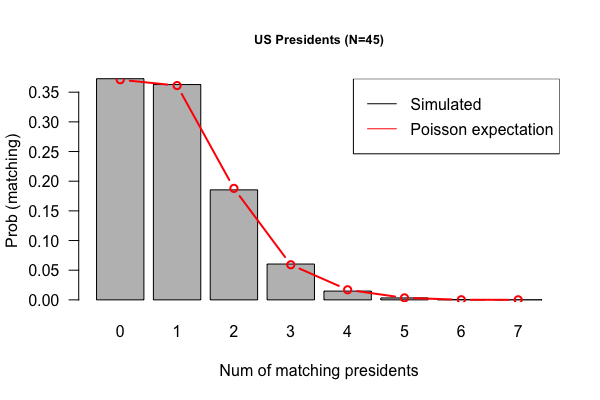
Feb 3, 2017 "US Presidents" (Due 2/10/2017)
|