Pseudomonas population genomics
Jump to navigation
Jump to search
Projects
- Build a local genome database
- Database schema:
- "genome": genome_id, strain_name, ncbi_taxid
- "orf": genome_id, locus_tag, start, stop, strand, genome_name, product_name
- "orth_orf": orth_orf_id, locus_name, genome_id, orth_class
- Parsing scripts
- Rayees Parsing code, requires that you remove columns 9-27 using bash command:
cut -c 1-8(I will write a bash script that does this and runs the program) https://www.dropbox.com/s/lpxxbkxeyw7frrn/parser.pl
- Rayees Parsing code, requires that you remove columns 9-27 using bash command:
- Database loading scripts
- Database schema:
- Molecular Evolution of flagellum genes
- Download orthologs
- Reconstruct phylogenetic tree
- Run PAML tests
Update: July 16, 2013
Treefiles based on aligned orthologs
| number of genes used | Method | treefile |
| 100 | FASTTREE | treefile |
| 100 | FASTTREE | Treefile 2 |
| all orthologs | Phylip | treefile_phylip |
FleN, FleQ, Flhf unique strains
| Gene | alignment | tree |
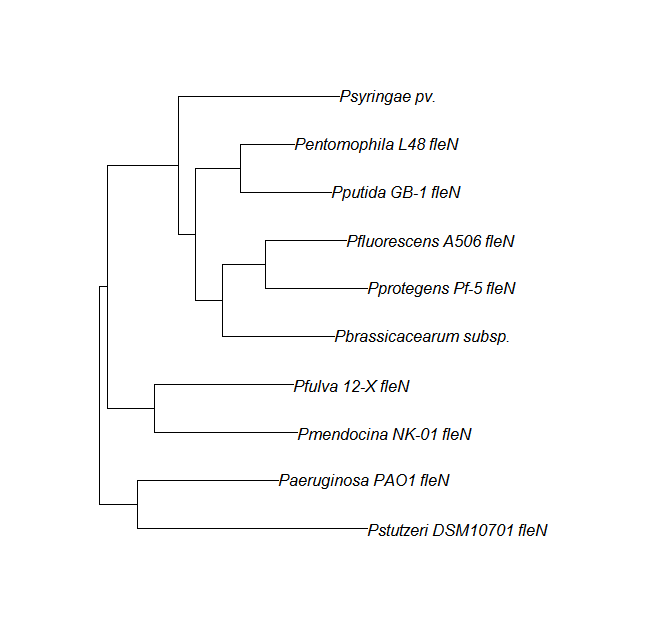
| fleN | fleN unique align | |
| fleQ | fleQ unique align | 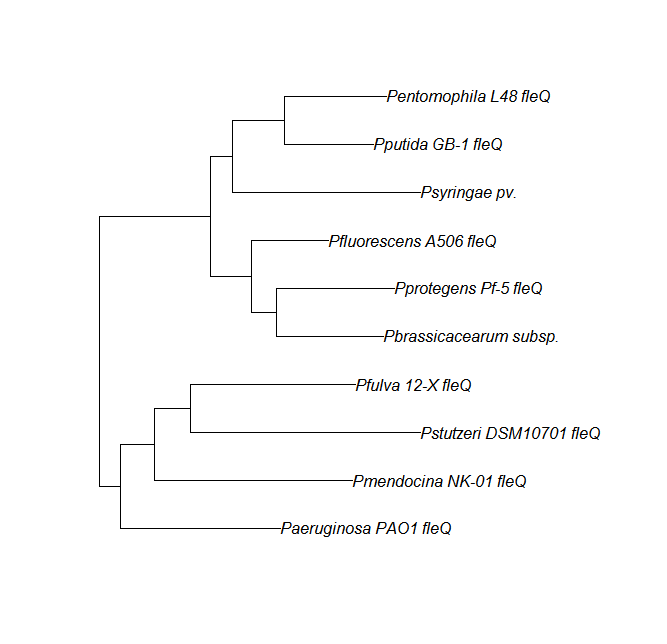
|
| flhf | flhF unique align | 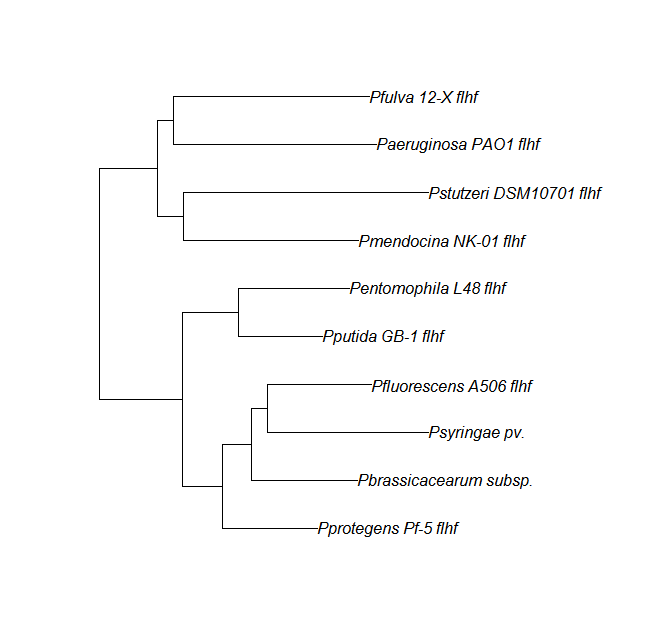
|
To do:
- Create pipeline
Update: June 28, 2013
Phylogenetic Analysis by Maximum Likelihood (PAML) Test performed on FleN, FleQ, FlhF orthologs and aeruginosa only orthologs
| Gene | PAML outfile for orthologs | |
|---|---|---|
| fleN | fleN PAML | |
| fleQ | fleQ PAML | |
| flhf | flhF-corrected PAML |
To do:
- Estimate genome tree using MrBayes and BEST (http://www.stat.osu.edu/~dkp/BEST/introduction/).
- Analysis of positively selected genes using PAML and homozygosity analysis
Update: June 18, 2013
Ortholog aligning and phylogenetic tree material and methods
Protein ortholog data: fleN, fleQ, flhF
- Protocol:
- Fasta headers are too long for tree, run: rename.pl to shorten names. Usage:
>rename.pl <FASTA_file> > <OUTPUTfilename.fas>(will rewrite script to create automatic output file) - To align use muscle Usage:
>muscle -in <FASTA_File> -out <OUTPUTfilename> -clwstrict - To create tree file run clustalW2 Usage
>clustalw2 -infile= <Aligned_file> -output= Phylip - To create tree run R using following commands:
setwd("<directory containing phylip files>")library ("ape")library ("phangorn")<gene_name> = read.tree("<file_gene_name.phy>")<gene_name> = midpoint(gene_name)plot(<gene_name>)
- Fasta headers are too long for tree, run: rename.pl to shorten names. Usage:
| Gene | Alignment | Tree | Notes |
|---|---|---|---|
| fleN | fleN pep alignment | 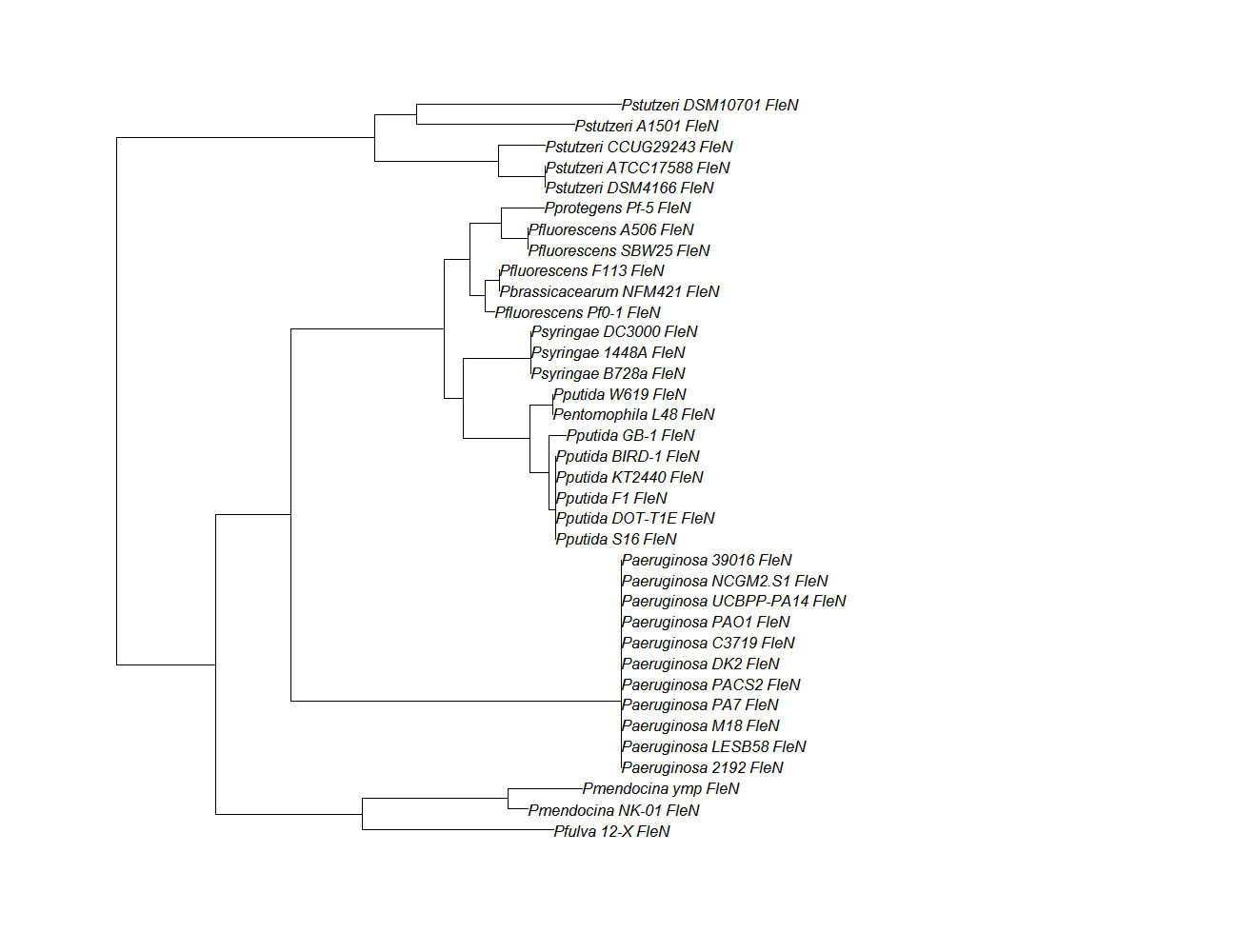 |
Conserved Domain |
| fleQ | fleQ pep alignment | 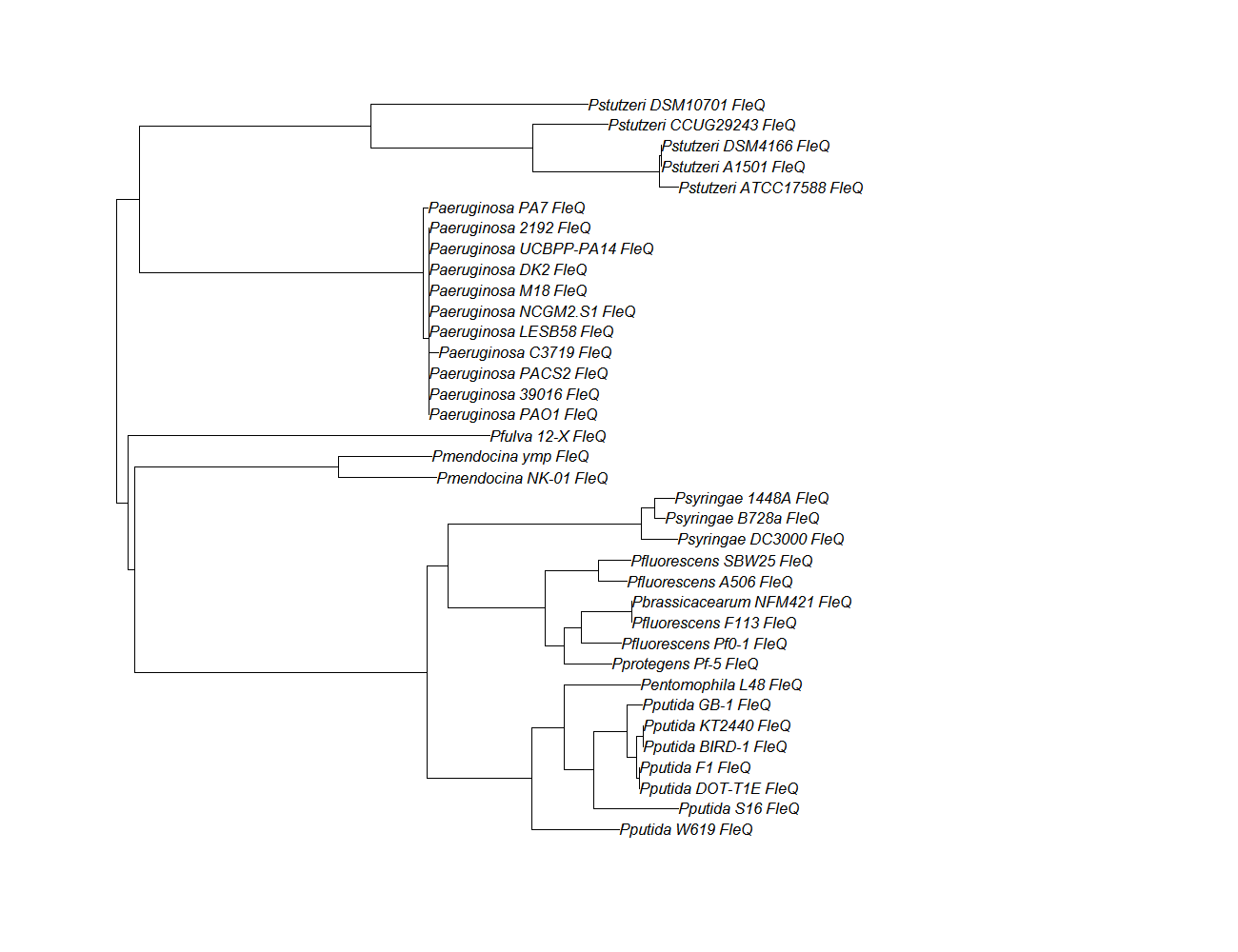 |
Conserved Domain |
| flhf | flhF pep alignment | 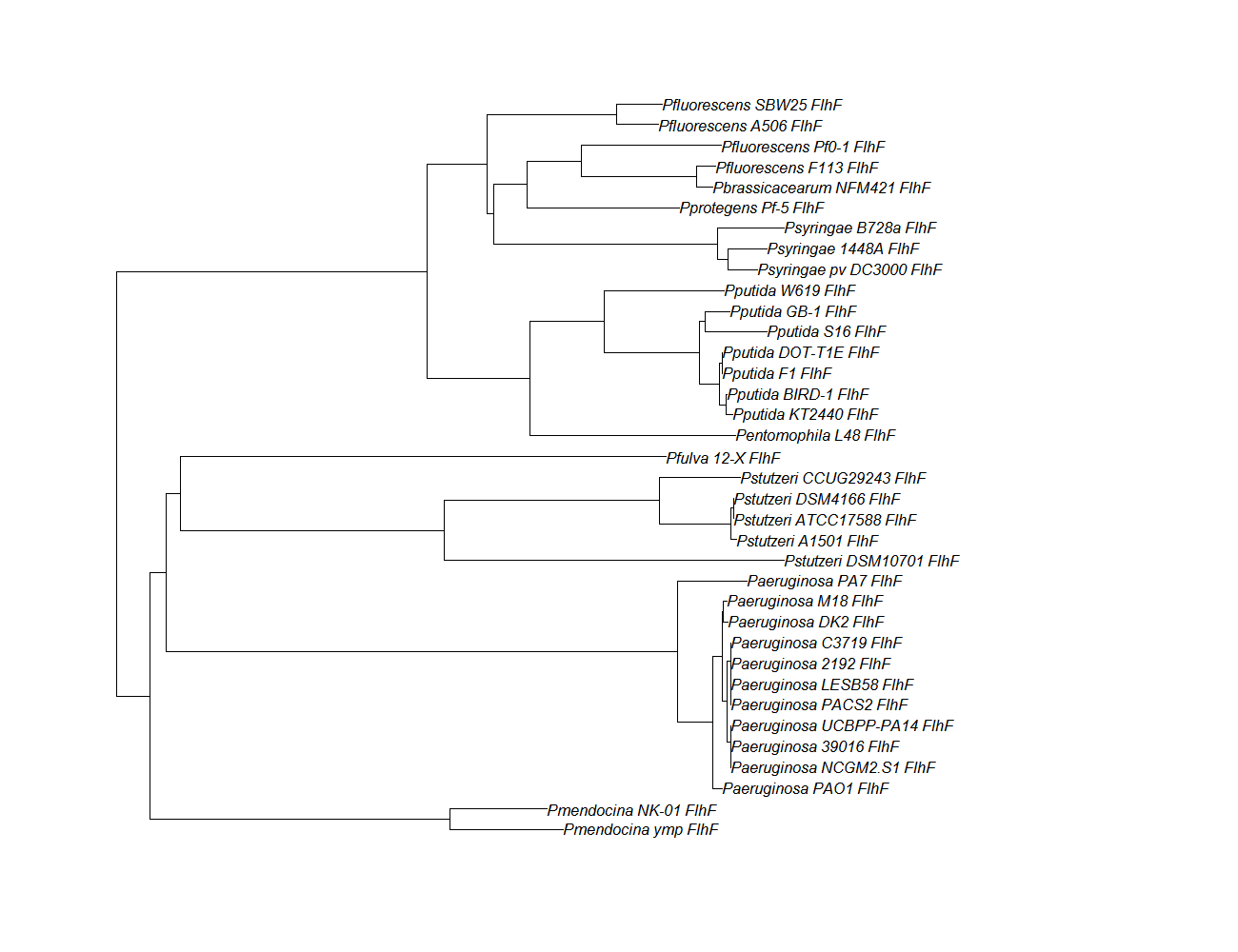 |
Conserved Domain |
Benchmark: June 11, 2013
- Finish parsing the genome files to upload the "orf" table (Raymond & Rayees)
- Rayees Parsed genome files: https://www.dropbox.com/sh/k0zktvvmv39op9i/1zBercEky8
- Parsing the ortholog file to upload the "orth_orf" table (Raymond)
- Identify and download fleN, fleQ, and flhF orthologs & align them (Rayees)