Main Page: Difference between revisions
Jump to navigation
Jump to search
m (→Web Apps) |
|||
| (48 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{| | {| | ||
|- | |- | ||
| [[File: | | | ||
[[File:Portrait-2022-blur.jpg| x250 px | thumb]] | |||
| <center> <strong>Welcome to Qiu Lab Wiki @ Hunter</strong><br>Weigang Qiu, Ph.D., Professor<br> [http://biology.hunter.cuny.edu Department of Biological Sciences]<br> | | <center> <strong>Welcome to Qiu Lab Wiki @ Hunter</strong><br>Weigang Qiu, Ph.D., Professor<br> [http://biology.hunter.cuny.edu Department of Biological Sciences]<br> | ||
[https://hunter.cuny.edu Hunter College] of<br> [https://www.cuny.edu City University of New York]<br> Belfer Research Building, Room 402<br>413 East 69th Street, New York, NY 10021<br>Office: 1-212-896-0445<br>Email: wqiu-at-(hunter.cuny.edu)<br>[https://goo.gl/maps/xv1KmaW3XEnxaY1V7 Directions by Google Map] | [https://hunter.cuny.edu Hunter College] of<br> [https://www.cuny.edu City University of New York]<br> Belfer Research Building, Room 402<br>413 East 69th Street, New York, NY 10021<br>Office: 1-212-896-0445<br>Email: wqiu-at-(hunter.cuny.edu)<br> </center> | ||
| [[File:Belfer_building.jpg | x300px | thumb | [https://goo.gl/maps/xv1KmaW3XEnxaY1V7 Directions by Google Map]]] | |||
| __TOC__ | |__TOC__ | ||
|} | |} | ||
| Line 11: | Line 12: | ||
===Lyme Genomics=== | ===Lyme Genomics=== | ||
<gallery mode="packed" heights="200px" perrow="4" style="text-align:left"> | <gallery mode="packed" heights="200px" perrow="4" style="text-align:left"> | ||
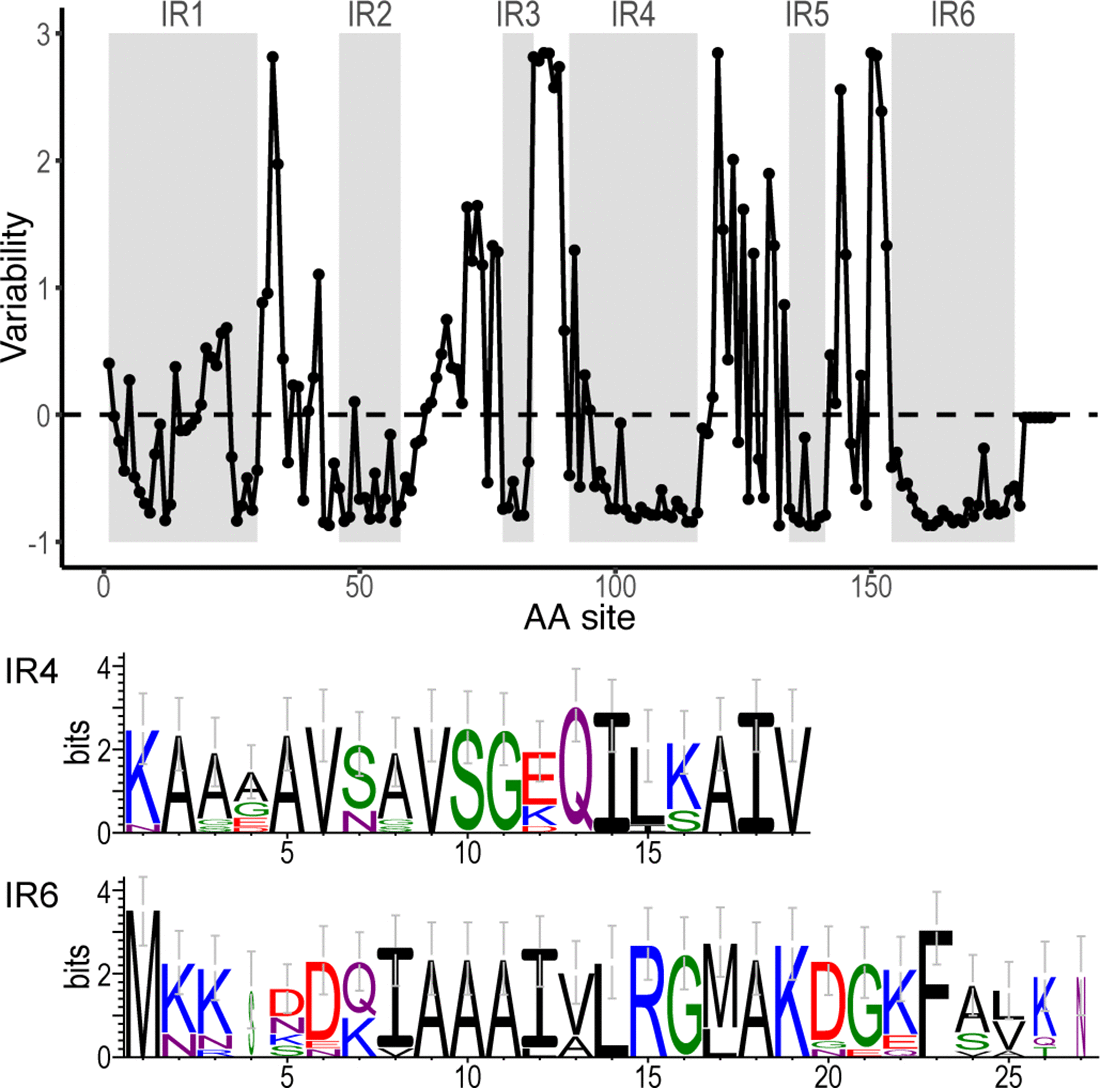
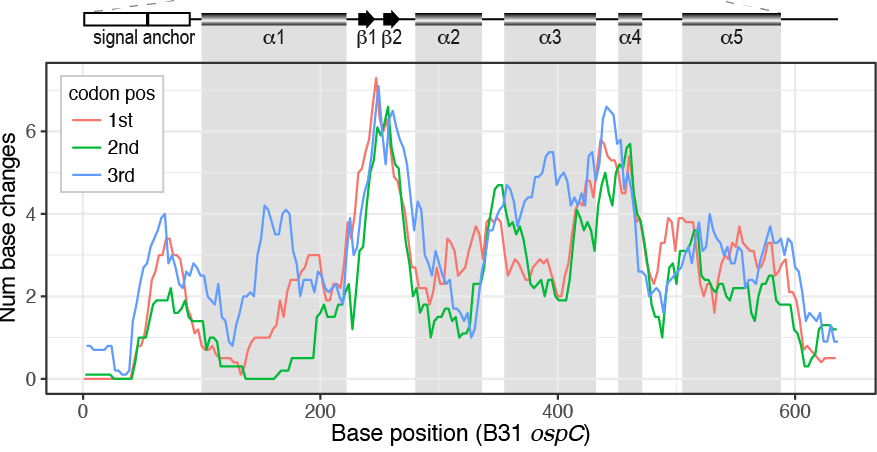
File:Spectrum.01743-22-f003.gif |Li, Di, Zeglis, Qiu (2022). “Evolution of the ''vls'' antigenic variability locus of the Lyme Disease pathogen and development of recombinant monoclonal antibodies targeting conserved VlsE epitopes”. '''''Microbial Spectrum''''' 10 (5):1-15. | File:Spectrum.01743-22-f003.gif | link=https://doi.org/10.1128/spectrum.01743-22 | Li, Di, Zeglis, Qiu (2022). “Evolution of the ''vls'' antigenic variability locus of the Lyme Disease pathogen and development of recombinant monoclonal antibodies targeting conserved VlsE epitopes”. '''''Microbial Spectrum''''' 10 (5):1-15. | ||
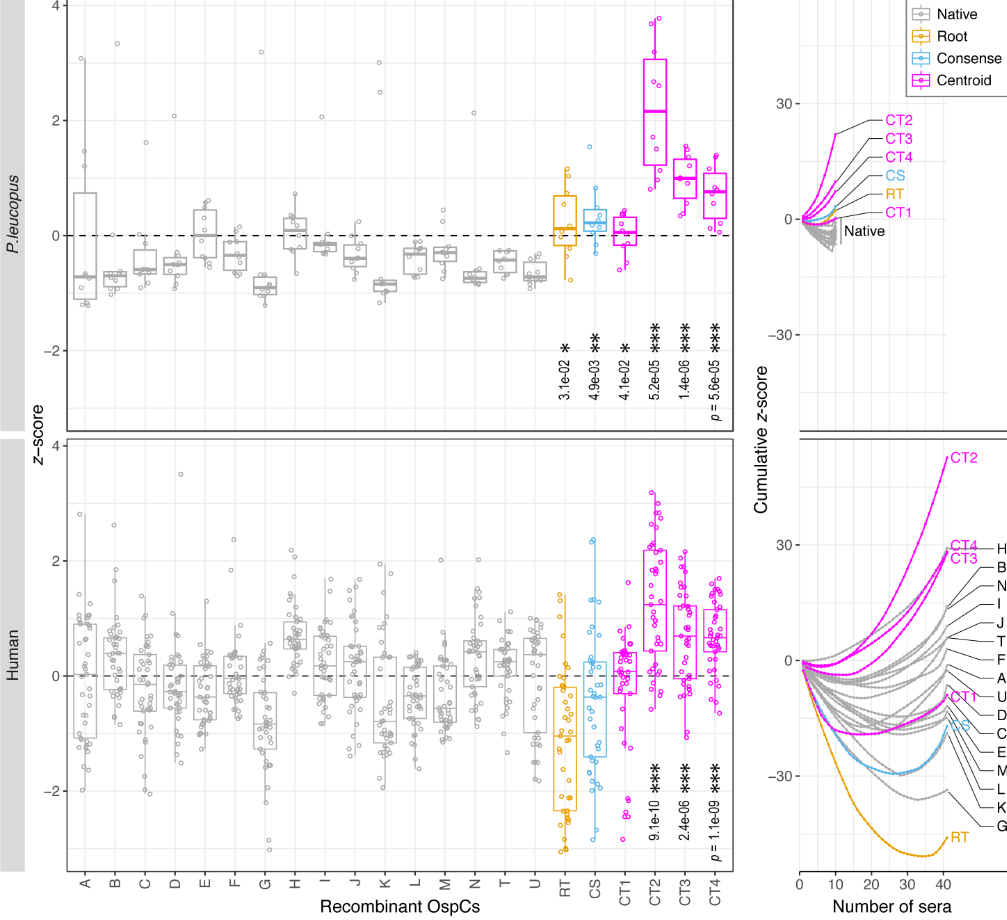
File:Fig7-small.png | link= https://pubmed.ncbi.nlm.nih.gov/34413477 | Di*, Akther*, Bezrucenkovas, Ivanova, Sulkow, Wu, Mneimneh, Gomes-Solecki, Qiu (2021). "Maximum antigen diversification in a lyme bacterial population and evolutionary strategies to overcome pathogen diversity". '''''The ISME Journal'''''. 16, 447-464. (*co-first authors) | |||
File:Ira-fig2.png|Schwartz, Margos, Casjens, Qiu, Eggers (2020). "Multipartite Genome of Lyme Disease ''Borrelia'': Structure, Variation and Prophages". '''''Current Issues in Molecular Biology'''''. 42:409-454. | File:Ira-fig2.png | link=https://doi.org/10.21775/9781913652616 | Schwartz, Margos, Casjens, Qiu, Eggers (2020). "Multipartite Genome of Lyme Disease ''Borrelia'': Structure, Variation and Prophages". '''''Current Issues in Molecular Biology'''''. 42:409-454. | ||
File:Fig6-1-Barbour.png | link=https://doi.org/10.1002/9781118960608.gbm01525 | Barbour & Qiu (2019). ''Borreliella''. In '''''Bergey's Manual of Systematics of Archaea and Bacteria'''''. John Wiley & Sons, Inc., in association with Bergey's Manual Trust. DOI: | |||
</gallery> | </gallery> | ||
===Evolution & Learning Algorithms=== | ===Evolution & Learning Algorithms=== | ||
<gallery mode="packed" heights="200px" perrow="3" style="text-align:left"> | <gallery mode="packed" heights="200px" perrow="3" style="text-align:left"> | ||
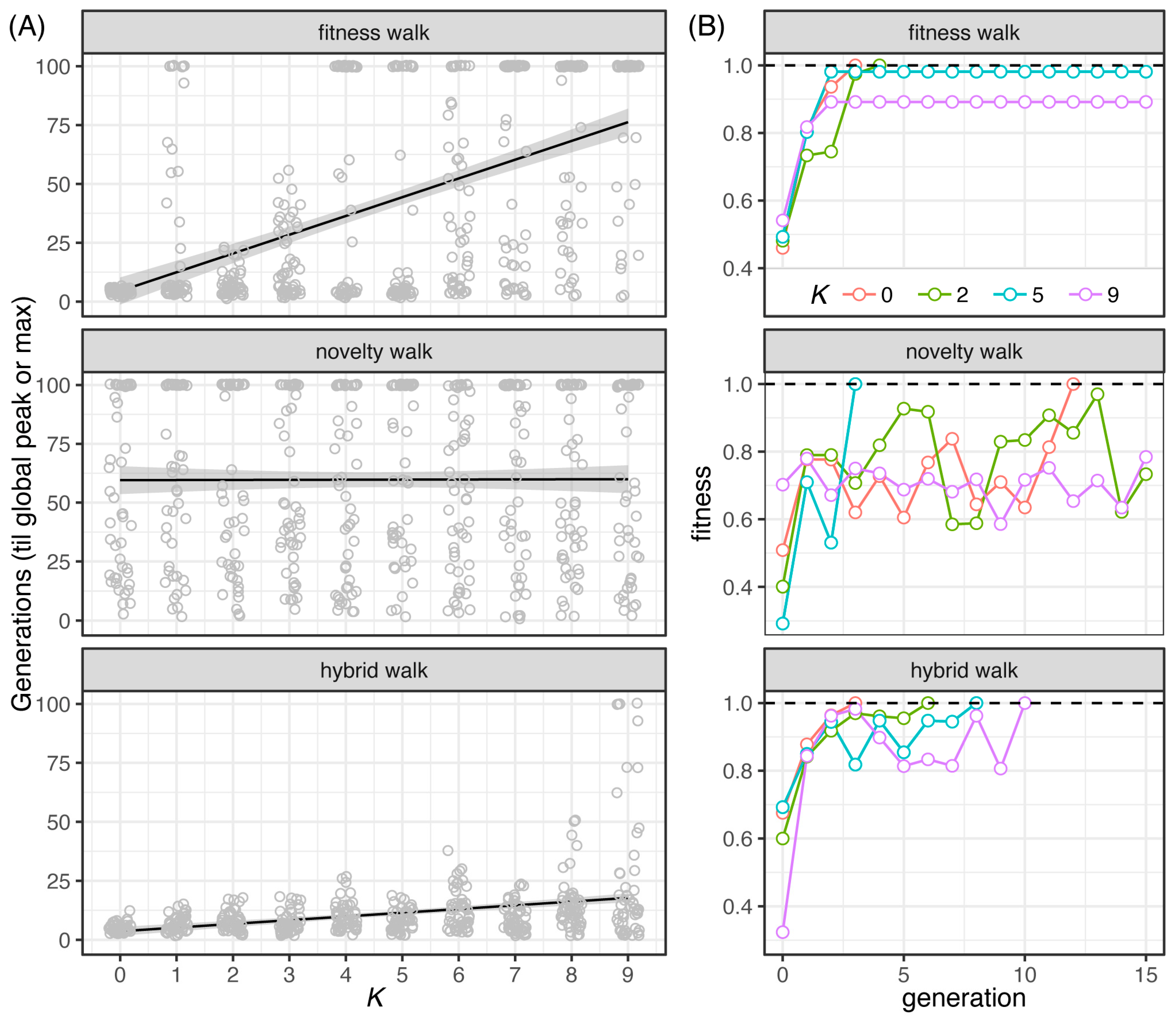
File:Pathogens-12-00388-g003.png|Ely, Koh, Ho, Hassan, Pham, Qiu (2023). Novelty Search Promotes Antigenic Diversity in Microbial Pathogens. '''''Pathogens''''' 12:388. | File:Pathogens-12-00388-g003.png | link=https://doi.org/10.3390/pathogens12030388 | Ely, Koh, Ho, Hassan, Pham, Qiu (2023). Novelty Search Promotes Antigenic Diversity in Microbial Pathogens. '''''Pathogens''''' 12:388. | ||
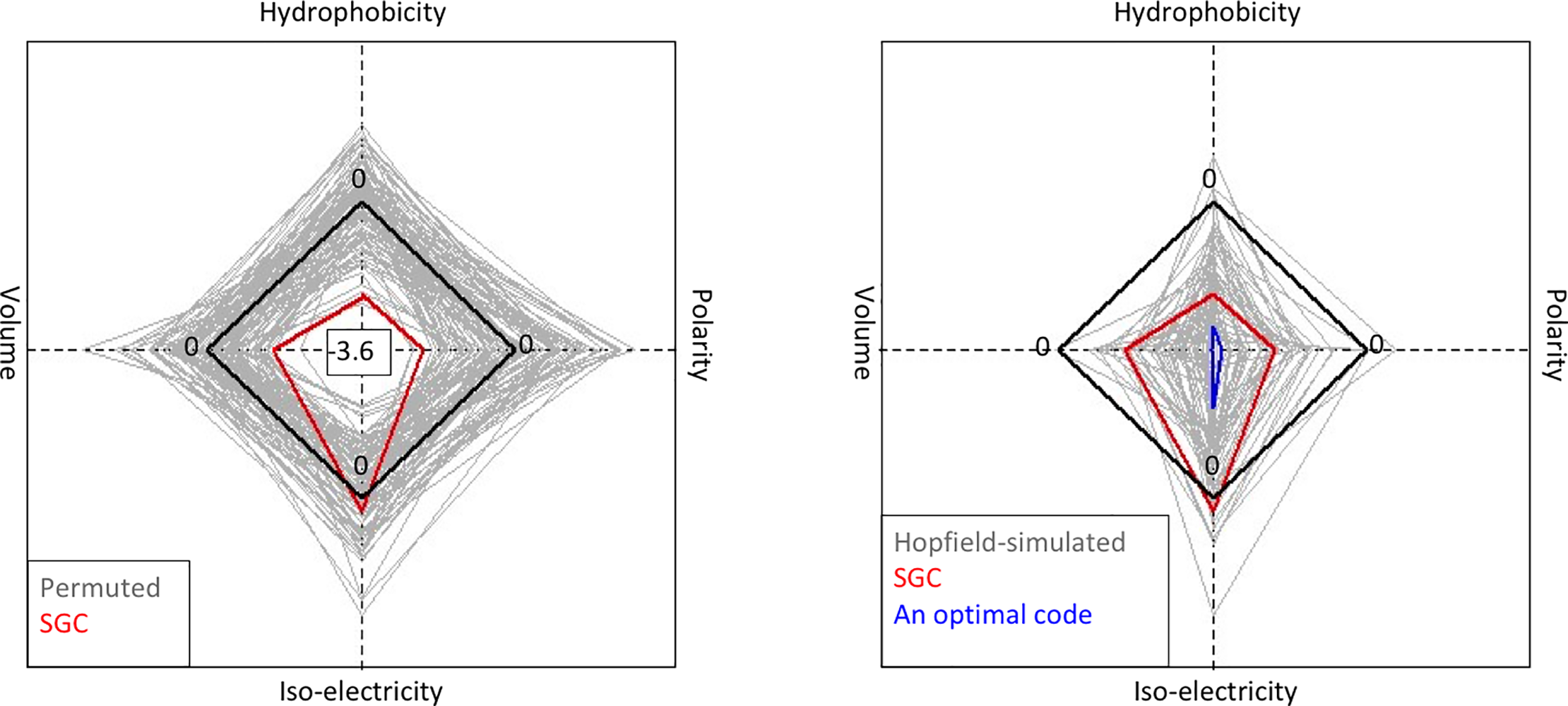
File:Pone.0224552.g005.png | link=https://doi.org/10.1371/journal.pone.0224552 | Attie, Sulkow, Di, Qiu (2019). Genetic codes optimized as a traveling salesman problem. '''''PLoS ONE''''' 14(10): e0224552. | |||
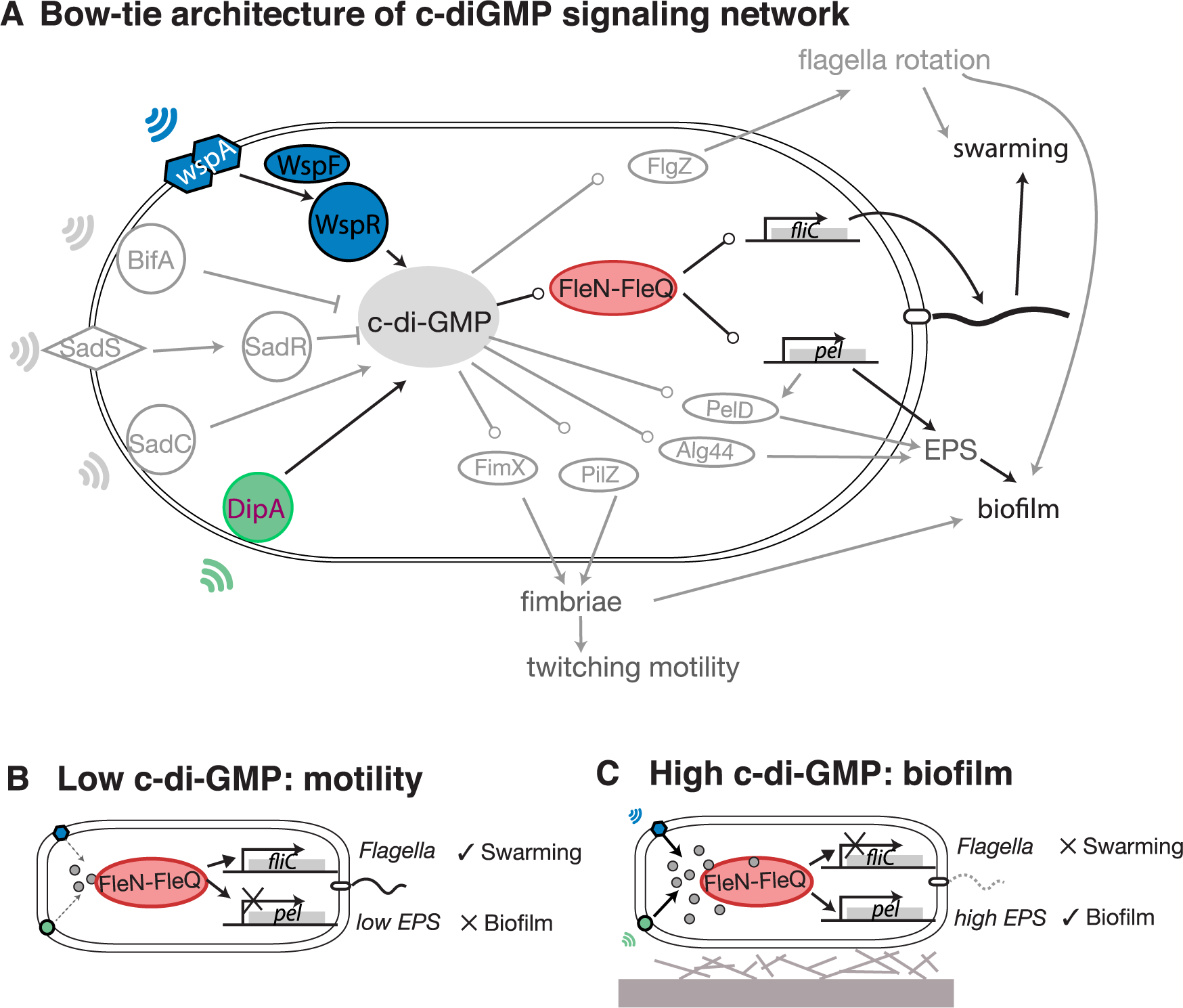
File:Pcbi.1005677.g001.png | link=https://doi.org/10.1371/journal.pcbi.1005677 | Yan, Deforet, Boyle, Rahman, Liang, Okegbe, Dietrich, Qiu, Xavier (2017). Bow-tie signaling in c-di-GMP: Machine learning in a simple biochemical network. '''''PLoS Comput Biol''''' 13(8): e1005677. | |||
</gallery> | </gallery> | ||
===Informatics Tool Development=== | ===Informatics Tool Development=== | ||
<gallery mode="packed" heights="200px" style="text-align:left"> | <gallery mode="packed" heights="200px" style="text-align:left"> | ||
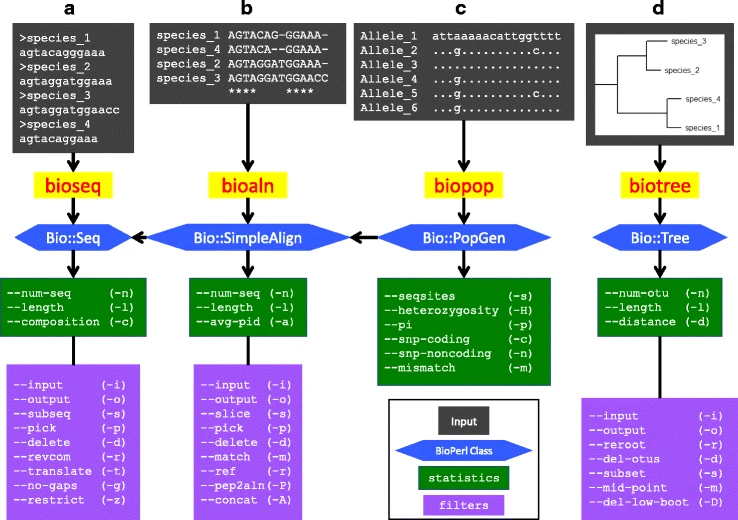
File: 12859 2018 2074 Fig1 HTML.png | Hernandez, Bernstein, Pagan, Vargas, McCaig, Ramrattan, Akther, Larracuente, Di, Vieira, Qiu. (2018). BbWrapper: BioPerl-based sequence and tree utilities for rapid prototyping of bioinformatics pipelines. '''''BMC Bioinformatics''''' 19(1):76. | File: 12859 2018 2074 Fig1 HTML.png | link=https://pubmed.ncbi.nlm.nih.gov/29499649 | Hernandez, Bernstein, Pagan, Vargas, McCaig, Ramrattan, Akther, Larracuente, Di, Vieira, Qiu. (2018). BbWrapper: BioPerl-based sequence and tree utilities for rapid prototyping of bioinformatics pipelines. '''''BMC Bioinformatics''''' 19(1):76. | ||
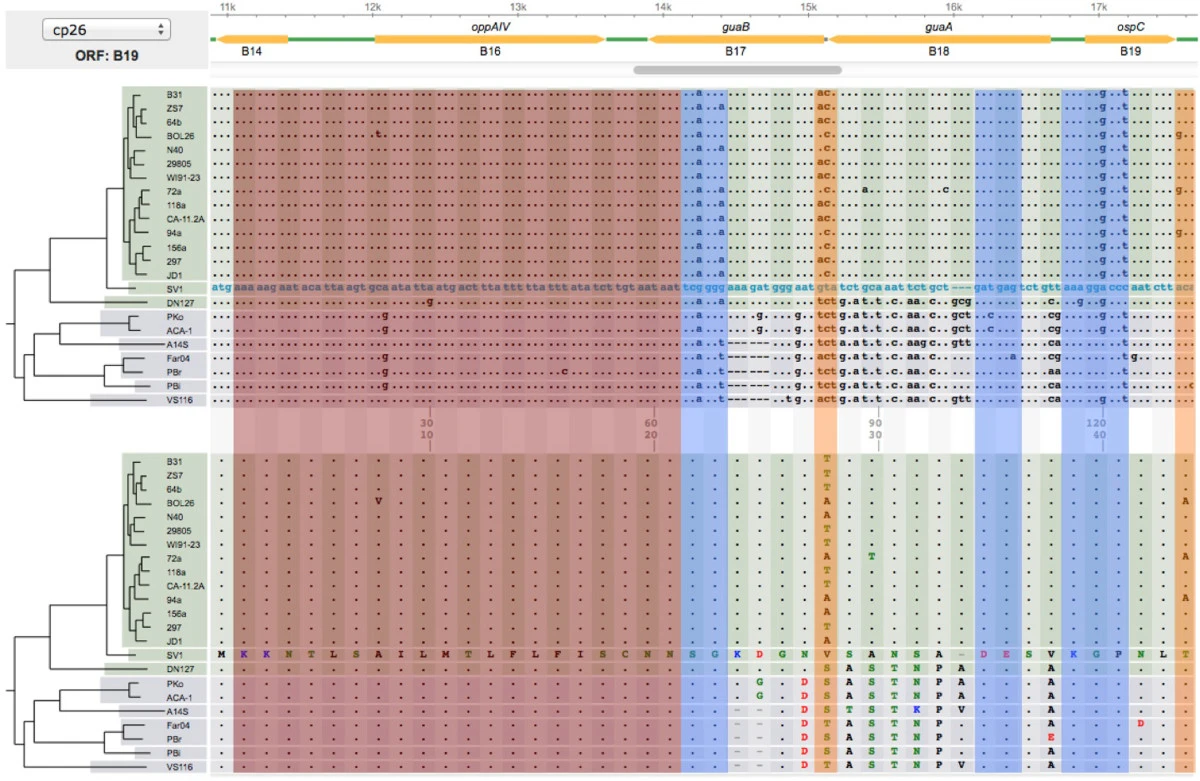
File: 12859 2014 Article 6488 Fig3 HTML.png | link=https://pubmed.ncbi.nlm.nih.gov/24994456 | Di, Pagan, Packer, Martin, Akther, Ramrattan, Mongodin, Fraser, Schutzer, Luft, Casjens and Qiu. (2014). BorreliaBase: a phylogeny-centered browser of ''Borrelia'' genomes. '''''BMC Bioinformatics''''' 15:233. | |||
</gallery> | </gallery> | ||
| Line 33: | Line 34: | ||
* [http://www.ncbi.nlm.nih.gov/sites/myncbi/weigang.qiu.1/bibliography/42770924/public/ Full list by NCBI Bibliography] | * [http://www.ncbi.nlm.nih.gov/sites/myncbi/weigang.qiu.1/bibliography/42770924/public/ Full list by NCBI Bibliography] | ||
last update: March 20, 2023 | last update: March 20, 2023 | ||
==Lab members and trainees== | ==Lab members and trainees== | ||
| Line 62: | Line 40: | ||
! Year/Period !! Doctoral members & trainees !! Other members & trainees | ! Year/Period !! Doctoral members & trainees !! Other members & trainees | ||
|- | |- | ||
| Current | | Current Academic Year | ||
(Fall 2022, Spring 2023 & Summer 2023) | |||
|| | || | ||
* Li Li (Lily): CUNY Grad Center, Biology/EEB doctoral program | * Li Li (Lily): CUNY Grad Center, Biology/EEB doctoral program | ||
| Line 78: | Line 54: | ||
|- | |- | ||
| Alumni | | Alumni (Since Fall 2002) | ||
|| | || | ||
* Dr Lia Di: Ph.D. from Nanjing Agricultural University & Wisconsin Blood Institute | * Dr Lia Di: Ph.D. from Nanjing Agricultural University & Wisconsin Blood Institute | ||
| Line 86: | Line 62: | ||
* Dr James Haven (2011): CUNY Grad Center, Biology/MCD | * Dr James Haven (2011): CUNY Grad Center, Biology/MCD | ||
* Dr Tika Sukarna (2009): CUNY Grad Center, Biology/MCD | * Dr Tika Sukarna (2009): CUNY Grad Center, Biology/MCD | ||
* Dr Juan Coronado (2008): CUNY Grad Center, Biology/MCD ( | * Dr Juan Coronado (2008): CUNY Grad Center, Biology/MCD (Dr Peter Lipke) | ||
* Dr William McCaig: CUNY BA, Ph.D. from Stony Brook University | * Dr William McCaig: CUNY BA, Ph.D. from Stony Brook University | ||
* Dr Vincent Xue: CUNY CS/Bioinformatics, Ph.D. from MIT | * Dr Vincent Xue: CUNY CS/Bioinformatics, Ph.D. from MIT | ||
* Dr Fubin Li: CUNY Grad Center, Biology/MCD (Dr Laurel Eckhardt) | |||
|| | || | ||
(published coauthors)<br> | (published coauthors)<br> | ||
| Line 94: | Line 71: | ||
* Winston Koh: Hunter Bio/CS | * Winston Koh: Hunter Bio/CS | ||
* Eamen Ho: Hunter Bio/Bioinformatics | * Eamen Ho: Hunter Bio/Bioinformatics | ||
* Ahn Pham: Hunter Bio/Bioinformatics | |||
* Chris Panlasigui: Hunter Bio/Bioinformatics | |||
* Amanda Amanda Larracuente: CUNY Grad Center, Biology/MCD | |||
* Pedro Pagan: Hunter Bio/Bioinformatics | * Pedro Pagan: Hunter Bio/Bioinformatics | ||
* Edgaras Bezrucenkovas: Hunter Chem/Bioinformatics | * Edgaras Bezrucenkovas: Hunter Chem/Bioinformatics | ||
| Line 102: | Line 82: | ||
* Mei Wu: CUNY City Tech | * Mei Wu: CUNY City Tech | ||
* Desiree Pante: Hunter Bio | * Desiree Pante: Hunter Bio | ||
* Saimtun Shipa: Hunter Stat (MA) | |||
* Bing Wu: Hunter Bio/Biotechnology | * Bing Wu: Hunter Bio/Biotechnology | ||
* Svidatoslav Kendall (Slav): Hunter Biology | * Svidatoslav Kendall (Slav): Hunter Biology | ||
* Philip Romov: Hunter CS | |||
|} | |} | ||
==Teaching & | ==Web Apps== | ||
* [[QuBi/modules/biol203-Lab4|BIOL20300 Molecular Genetics, Lab 4]] | ===Apps with Collaborators=== | ||
* [[QuBi/modules/biol203-Lab7|BIOL20300 Molecular Genetics, Lab 7]] | <gallery mode="packed" heights="200px" perrow="4" style="text-align:left"> | ||
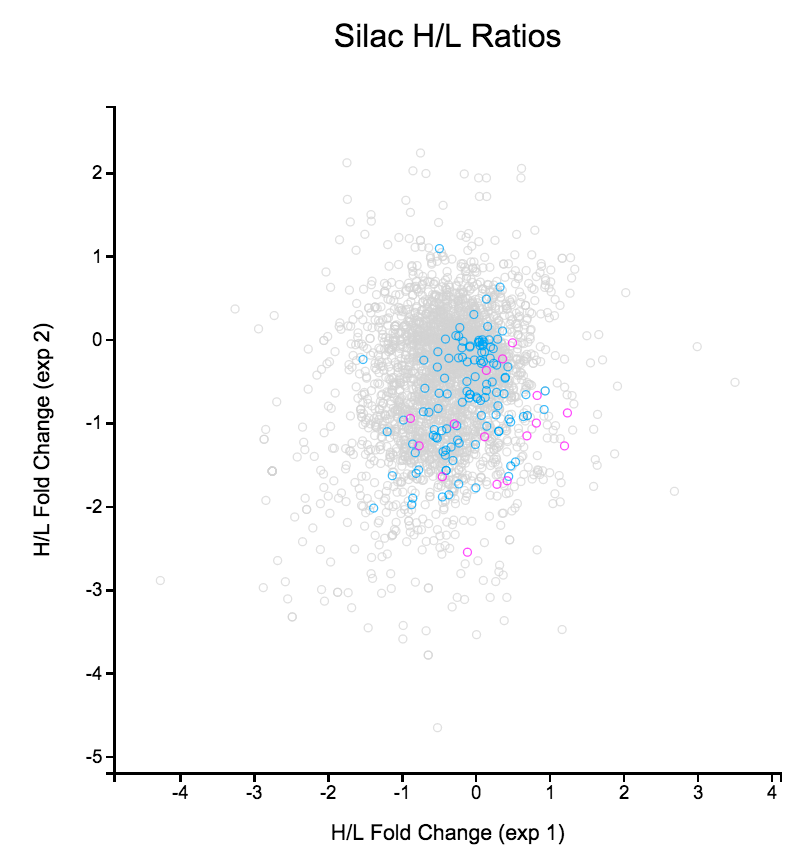
* [[QuBi/modules/biol303|BIOL30300 Cell Biology, Bioinformatics Lab (transcriptome analysis)]] | File: Silac.png | link=http://borreliabase.org/~wgqiu/Polotskaia_etal_2014/supp-fig-s1/ | Data & [http://borreliabase.org/~wgqiu/Polotskaia_etal_2014/supp-table-s1/ GSEA] associated with Polotskaia_etal_2014 (with Bargonetti Lab @Hunter) | ||
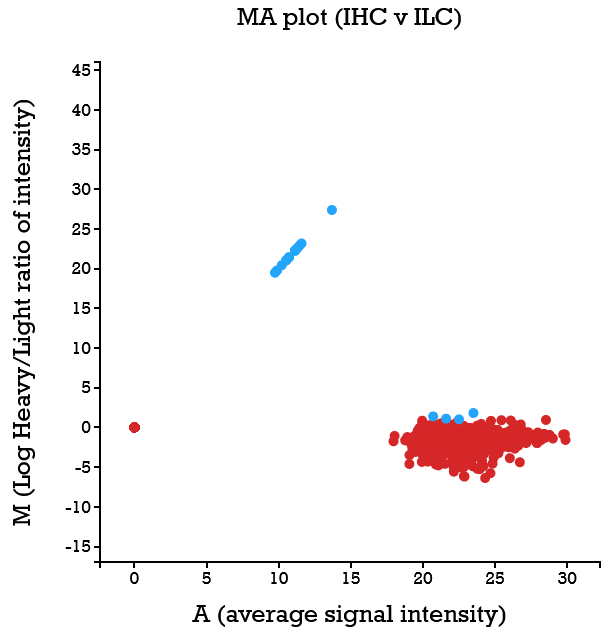
File:Screenshot_2023-03-24_122016.png | link=http://borreliabase.org/~wgqiu/clickme-khalikuz/temp-Points.html | Phosphoproteome MDM2KD data set (with Bargonetti Lab @Hunter) | |||
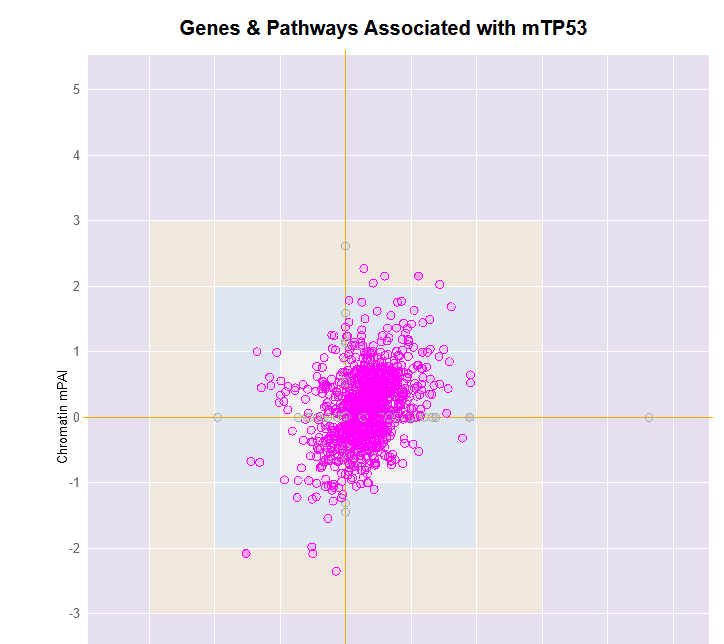
File:Screenshot_2023-03-24_122312.png | link=http://borreliabase.org/~wgqiu/mpai-v3/ | Genes & Pathways Associated with mTP53 (with Bargonetti Lab @Hunter) | |||
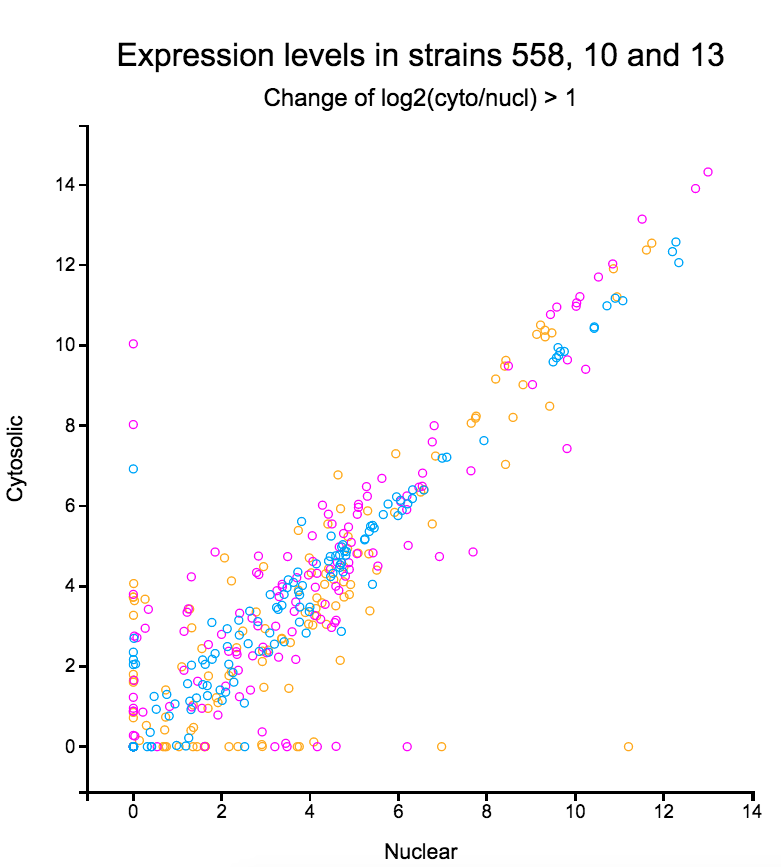
File:Spombe.png | link=Spombe | S. pombe transcriptomes (with Zhong Lab @Hunter) | |||
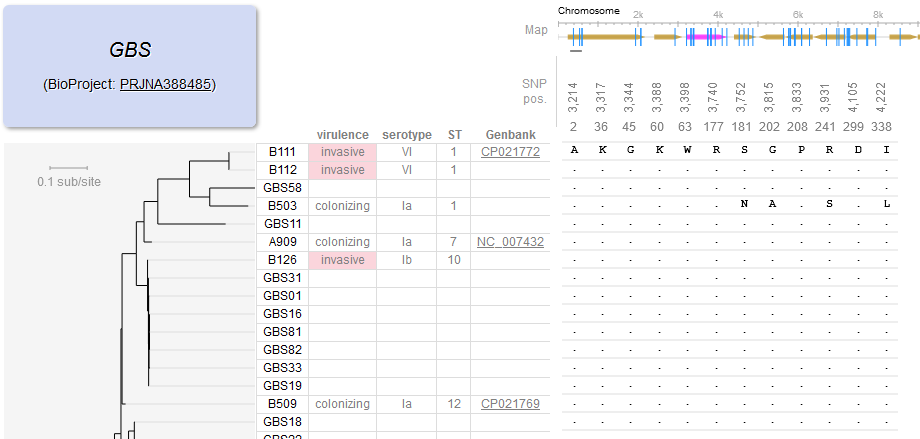
File:Screenshot 2023-03-24 122223.png | link=http://borreliabase.org/~wgqiu/gbs-browser-v3/ | Group B Streptococcus (GBS) genome browser (with Wu Lab @Shenzhen) | |||
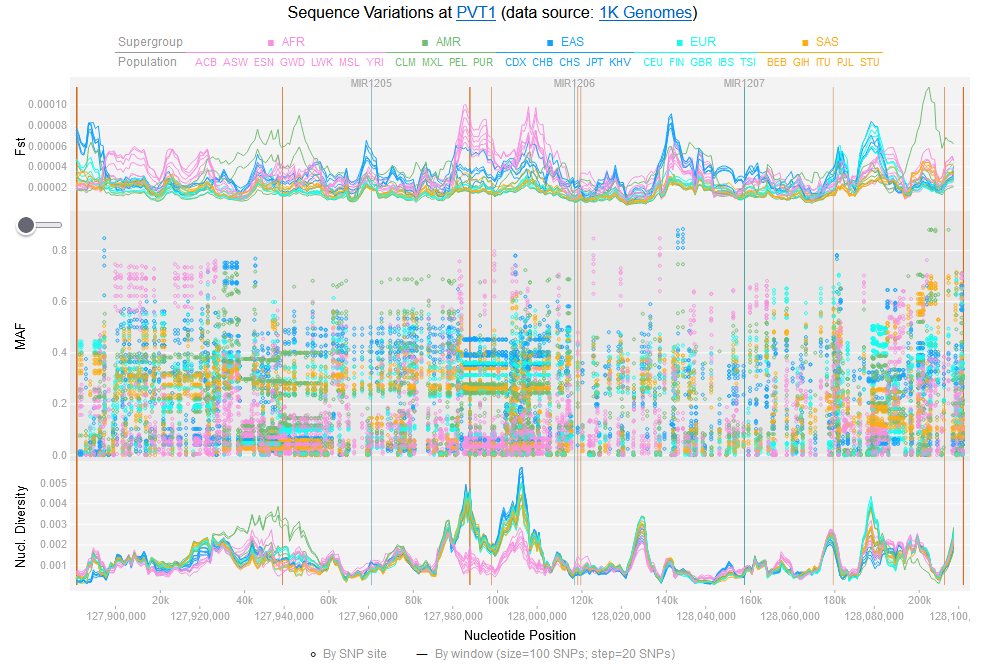
File:Screenshot 2023-03-24 122245.png | link=http://borreliabase.org/~wgqiu/oneKGenome/ | 1K genome (with Ogunwobi Lab @Hunter) | |||
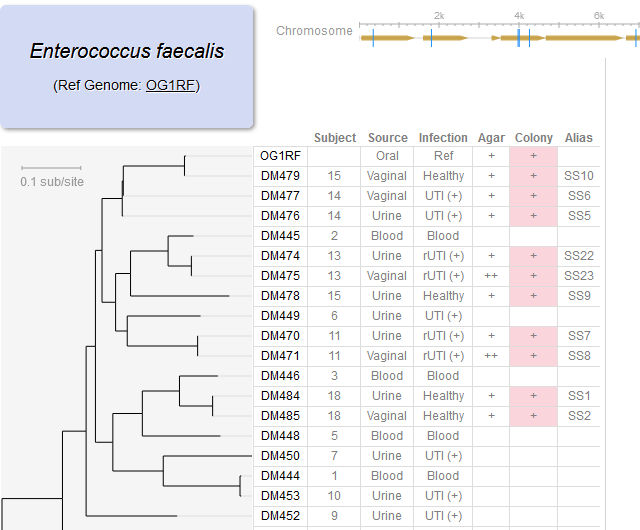
File:Screenshot 2023-03-24 122450.png | link=http://borreliabase.org/~wgqiu/E_faecalis/ | E faecalis genome browser (with Morales Lab @WCMC) | |||
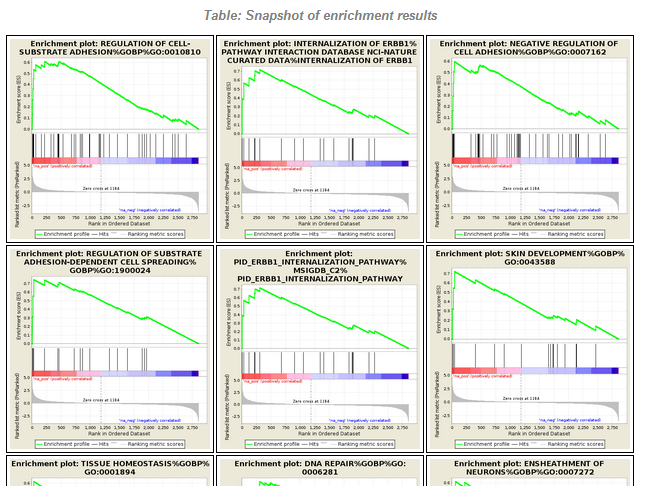
File:Screenshot 2023-03-24 122145.png | link=http://borreliabase.org/~wgqiu/carmen-proteomics/ | Proteomics GSEA results (with Melendez Lab @Hunter) | |||
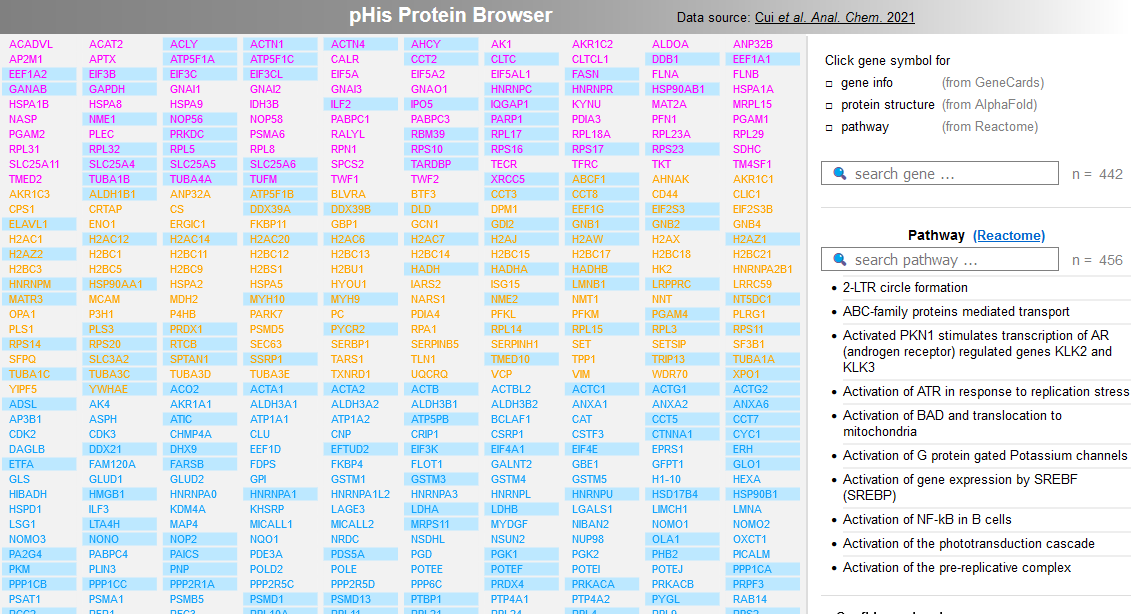
File:Screenshot 2023-03-24 122051.png | link=http://borreliabase.org/~wgqiu/Tcell-pHis-ACS/ | pHis Protein Browser (with Skolnik lab @NYU) | |||
</gallery> | |||
===Qiu Lab Apps=== | |||
<gallery mode="packed" heights="200px" perrow="3" style="text-align:left"> | |||
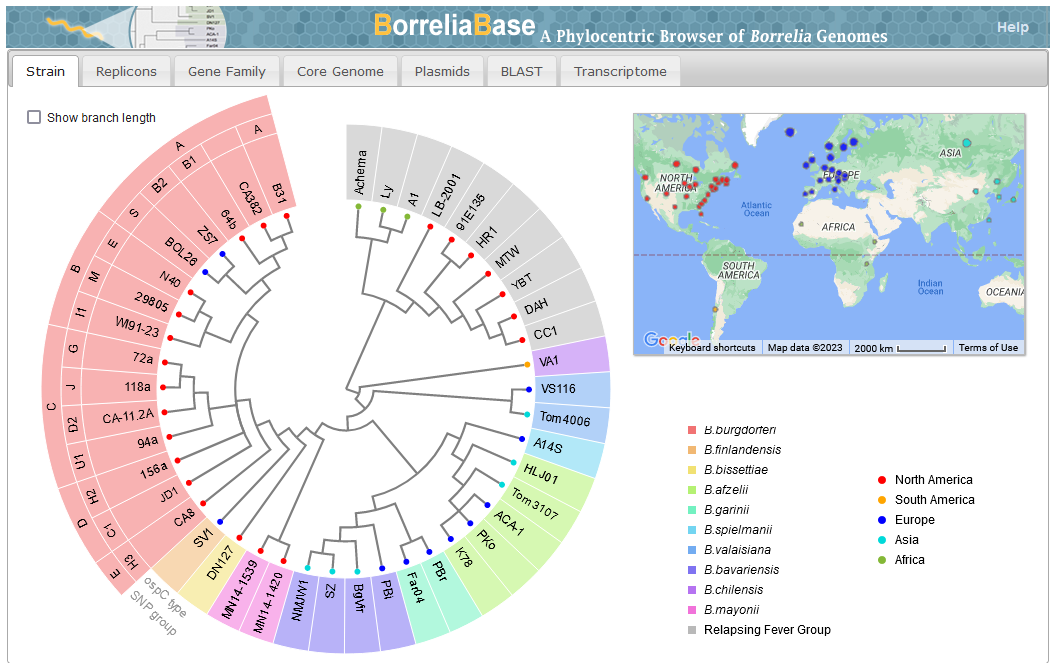
File:Screenshot 2023-03-24 122558.png | link=http://borreliabase.org | Lyme genome browser | |||
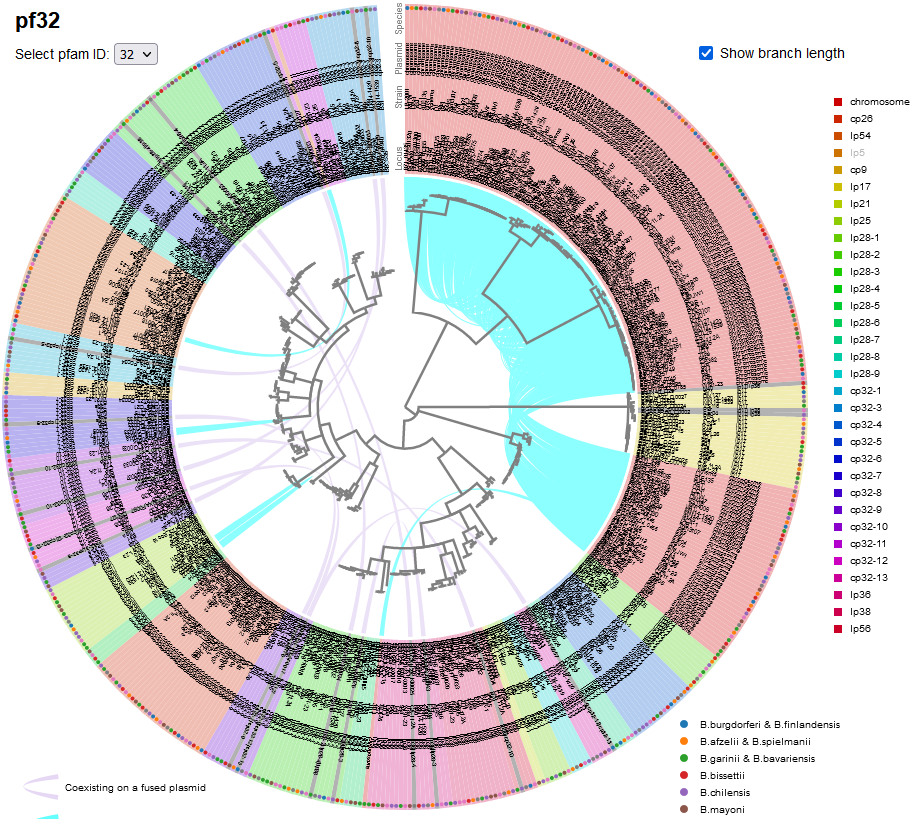
File:Screenshot 2023-03-24 133320.png | link=http://borreliabase.org/~wgqiu/pf-trees/ | Lyme pathogen plasmid partitioning gene trees | |||
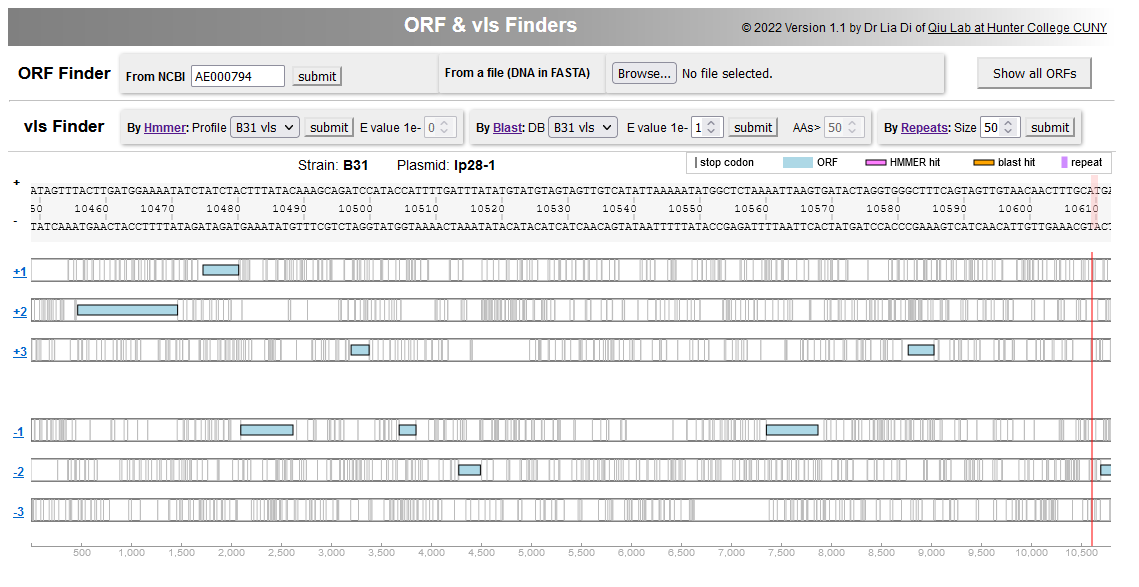
File:Screenshot 2023-03-24 133636.png | link=http://borreliabase.org/vls-finder/ | vls Finder in Lyme pathogen genomes | |||
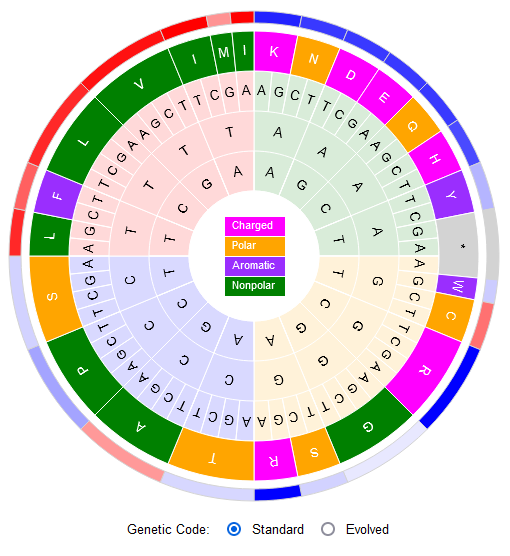
File:Screenshot 2023-03-24 122359.png | link=http://borreliabase.org/~wgqiu/code-wheel | Codon Wheel | |||
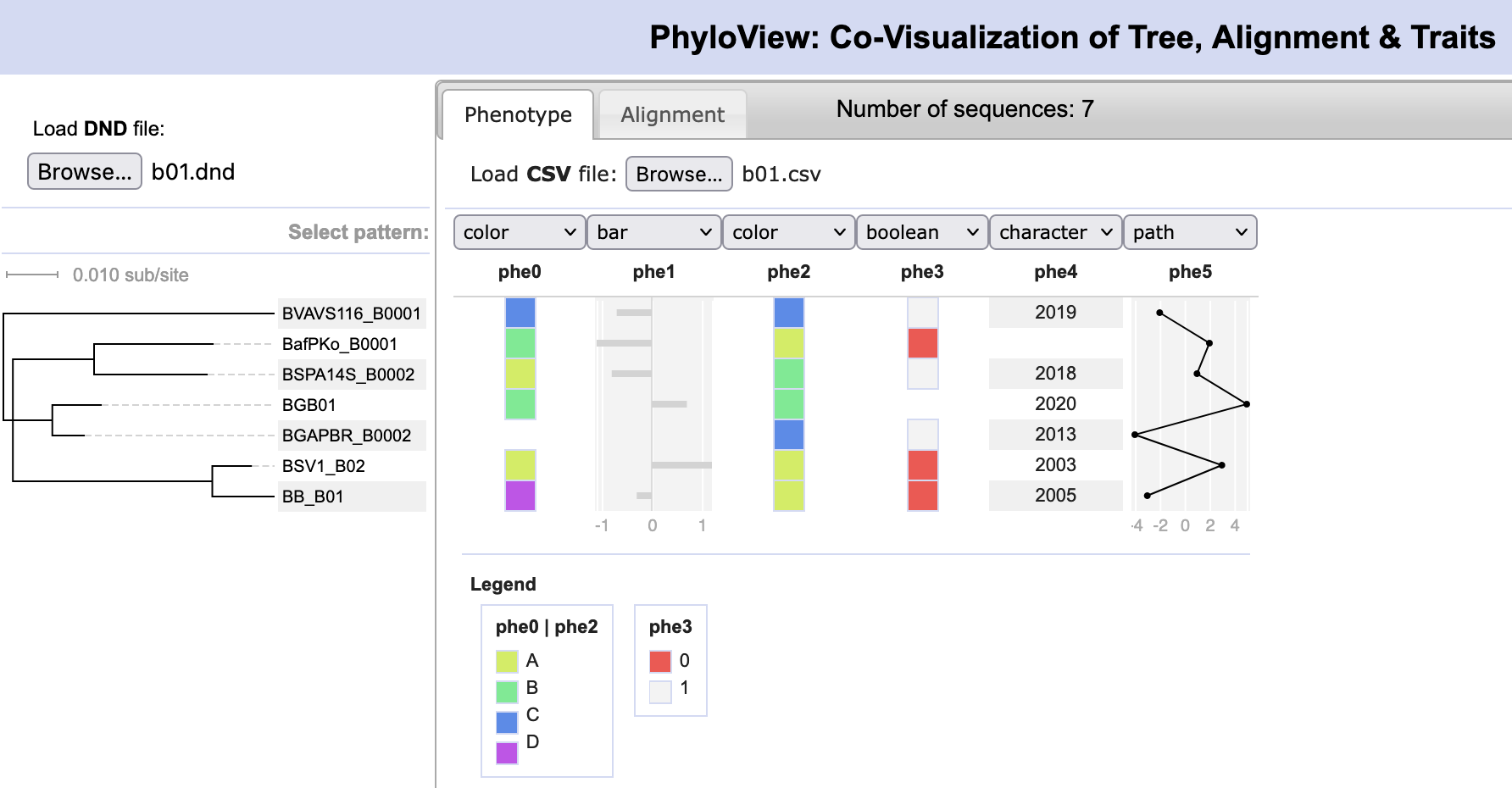
File:PhyloView.png | link=http://borreliabase.org/~wgqiu/PhyloView | Co-visualization of a tree with an alignment and character matrix | |||
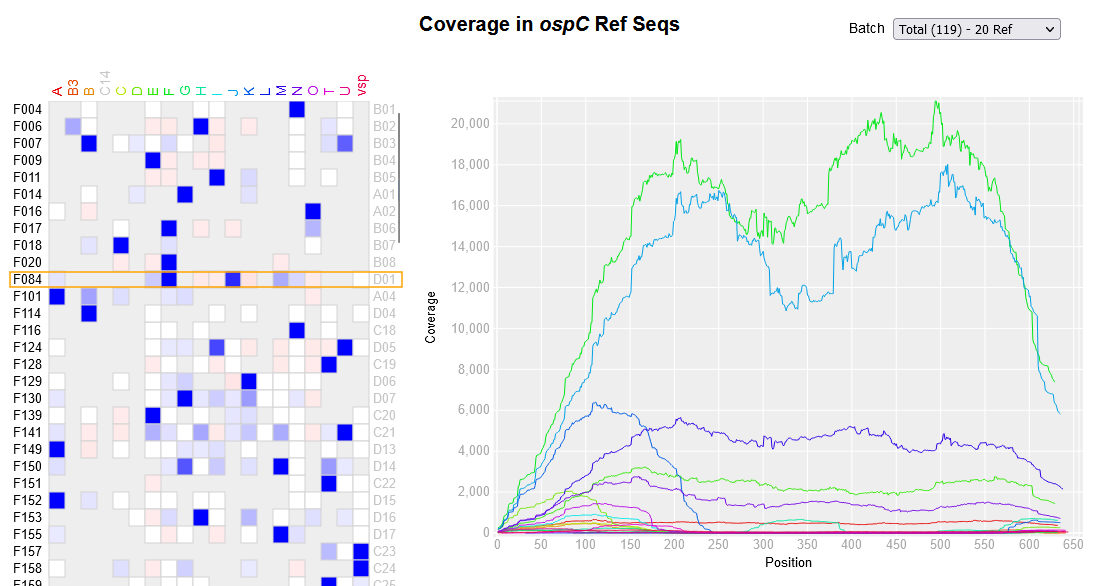
File:Screenshot 2023-03-24 122521.png | link=http://borreliabase.org/~wgqiu/ospC-sequencing | OspC amplicon sequencing from ticks | |||
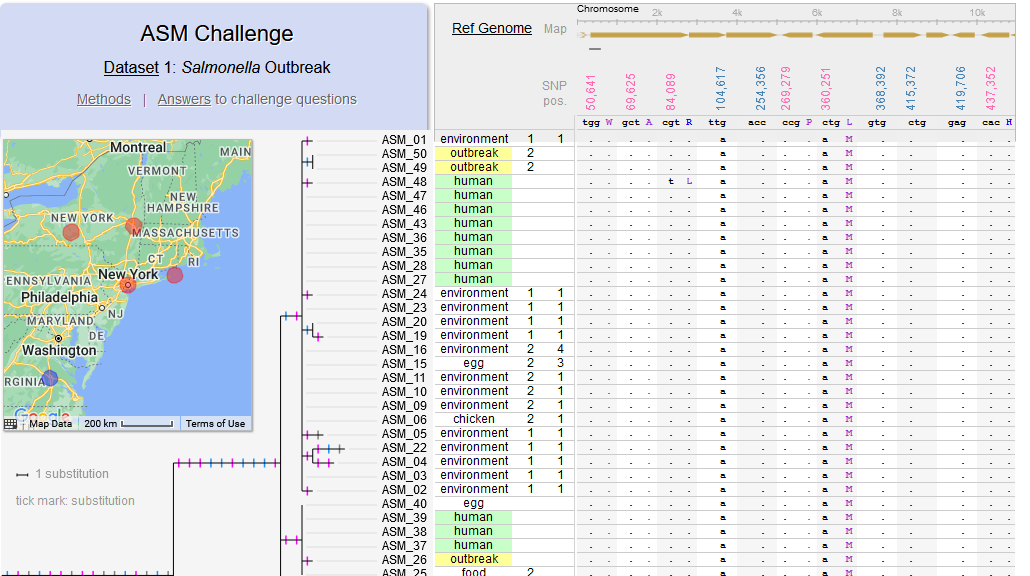
File:Screenshot 2023-03-24 122340.png | link=http://borreliabase.org/~wgqiu/asm-challenge | Genomic epidemiology of a Salmonella outbreak (ASM Challenge) | |||
</gallery> | |||
==Teaching & Curricular Development== | |||
* The Quantitative Biology (QuBi) Program at Hunter | |||
** [https://hunter-undergraduate.catalog.cuny.edu/programs/BIO1-BA Hunter Biology Major I (including the Bioinformatics Conentration)] | |||
** [https://hunter-undergraduate.catalog.cuny.edu/programs/CHEM2-BA Hunter Chemistry Major 2 (including the Bioinformatics Option)] | |||
** [https://hunter-undergraduate.catalog.cuny.edu/programs/COMPSCI-BA Hunter Computer Science (including Bioinformatics Concentration)] | |||
** [https://hunter-undergraduate.catalog.cuny.edu/programs/MATH-BA Hunter Mathematics (including the Quantitative Biology Concentration)] | |||
** [https://hunter-undergraduate.catalog.cuny.edu/programs/STATS-BA Hunter Statistics (including the Quantitative Biology Concentration)] | |||
** QuBi module: [[QuBi/modules/biol203-Lab4|BIOL20300 Molecular Genetics, Lab 4]] | |||
** QuBi module: [[QuBi/modules/biol203-Lab7|BIOL20300 Molecular Genetics, Lab 7]] | |||
** QuBi module: [[QuBi/modules/biol303|BIOL30300 Cell Biology, Bioinformatics Lab (transcriptome analysis)]] | |||
* [[BigData 2020|Big Data (2020)]] | * [[BigData 2020|Big Data (2020)]] | ||
* [[Southwest-University|Southwest University R course (2019]] | * [[Southwest-University|Southwest University R course (2019)]] | ||
* [[Biol425 2020|BIOL425 Computational Molecular Biology (2020)]] | * [[Biol425 2020|BIOL425 Computational Molecular Biology (2020)]] | ||
* [[Biol20N02 2017|Analysis of Biological Data (2017)]] | * [[Biol20N02 2017|Analysis of Biological Data (2017)]] | ||
* [[Biol375 2019|BIOL37500, Molecular Evolution (2019)]] | * [[Biol375 2019|BIOL37500, Molecular Evolution (2019)]] | ||
* [[Bioinformatics_Workshop_2014|Bioinformatics Workshop (2014)]] | |||
* [[BioMed-R-2020|BIOL47120 Biomedical Genomics II (2020)]] | |||
==SARS-CoV-2 genome evolution== | |||
<gallery mode="packed" heights="200px" perrow="3" style="text-align:left"> | |||
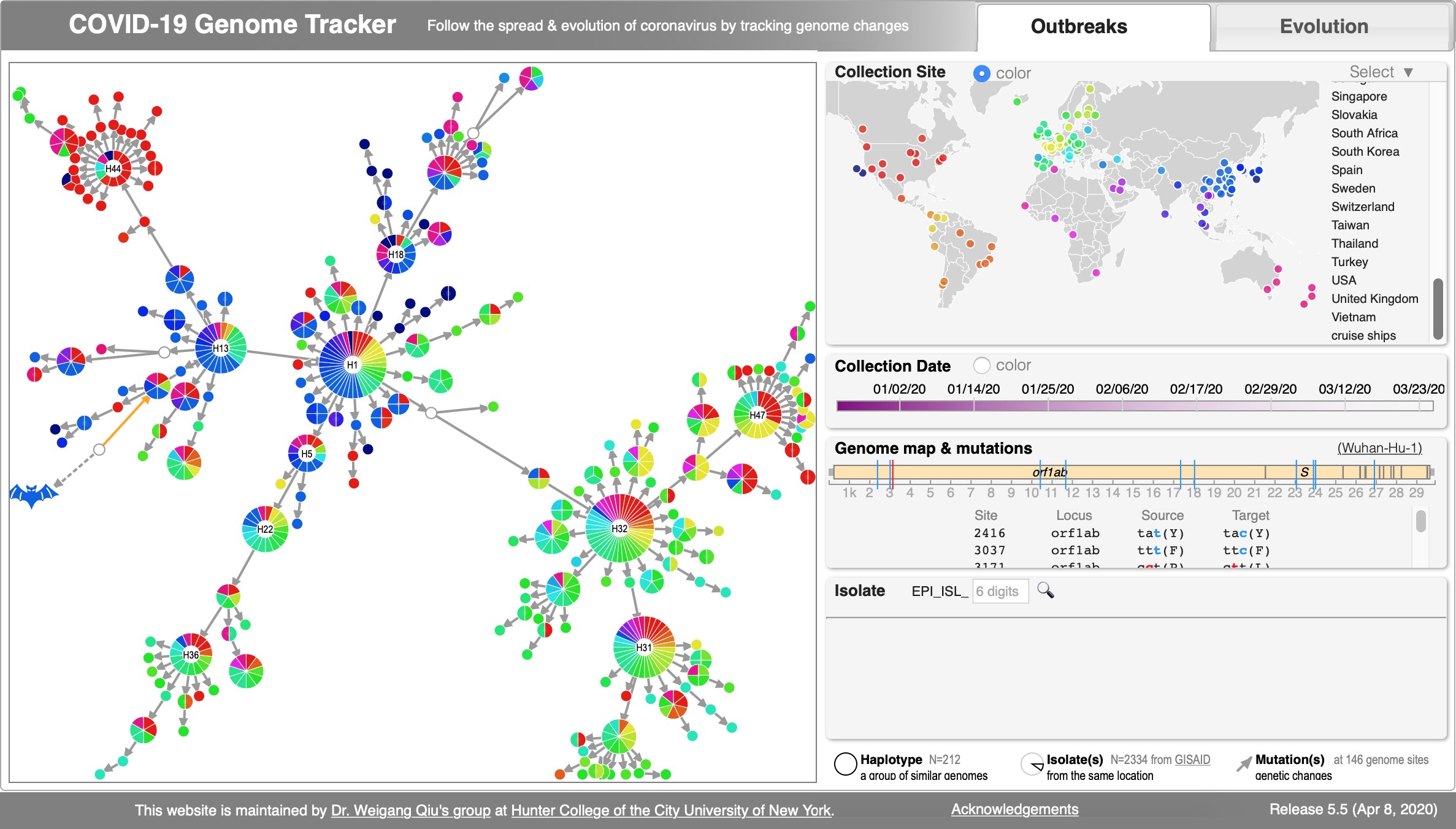
File:Cov-fig1.jpg | Akther, Bezrucenkovas, Sulkow, Panlasigui, Qiu, Di (April, 2020). "CoV Genome Tracker: tracing genomic footprints of Covid-19 pandemic". '''''[https://www.biorxiv.org/content/biorxiv/early/2020/04/14/2020.04.10.036343.full.pdf BioRxiv]'''''; Web app: SARS-CoV-2 Genome Tracker; Github: https://github.com/weigangq/cov-browser | |||
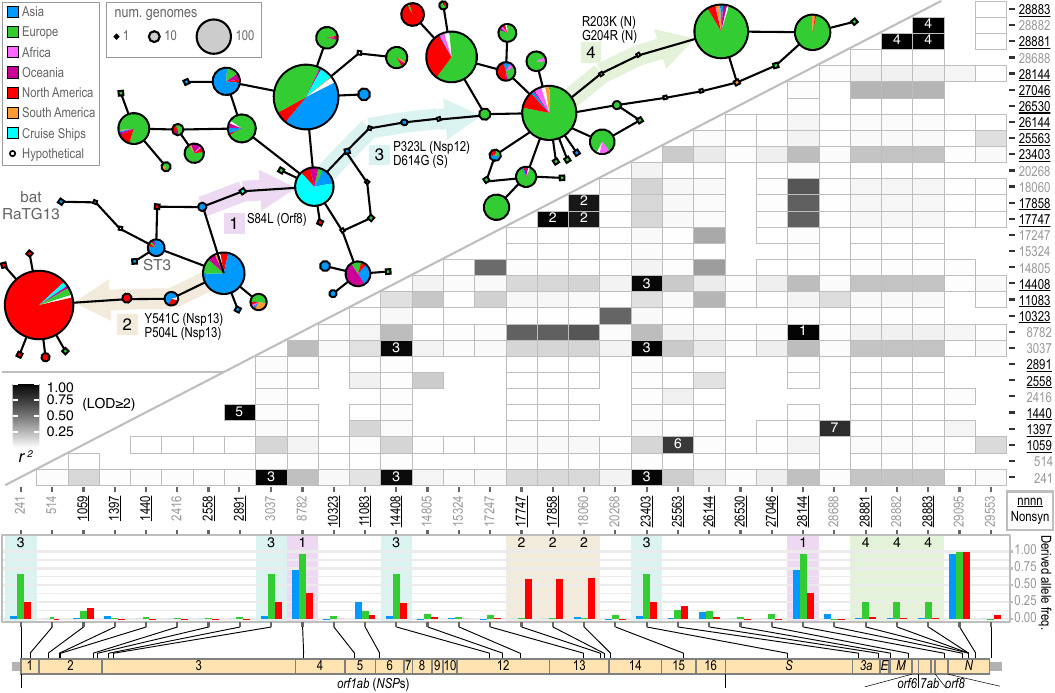
File:Rec-fig2.png | Akther, Li, Martin, Di, Sulkow, Pante, Bezrucenlovas, Luft, Qiu (May, 2020). "Origin, recombination, and missed opprotunities: a genomic perspective of the first 100 days of COVID-19 pandemic". (Unpublished). | |||
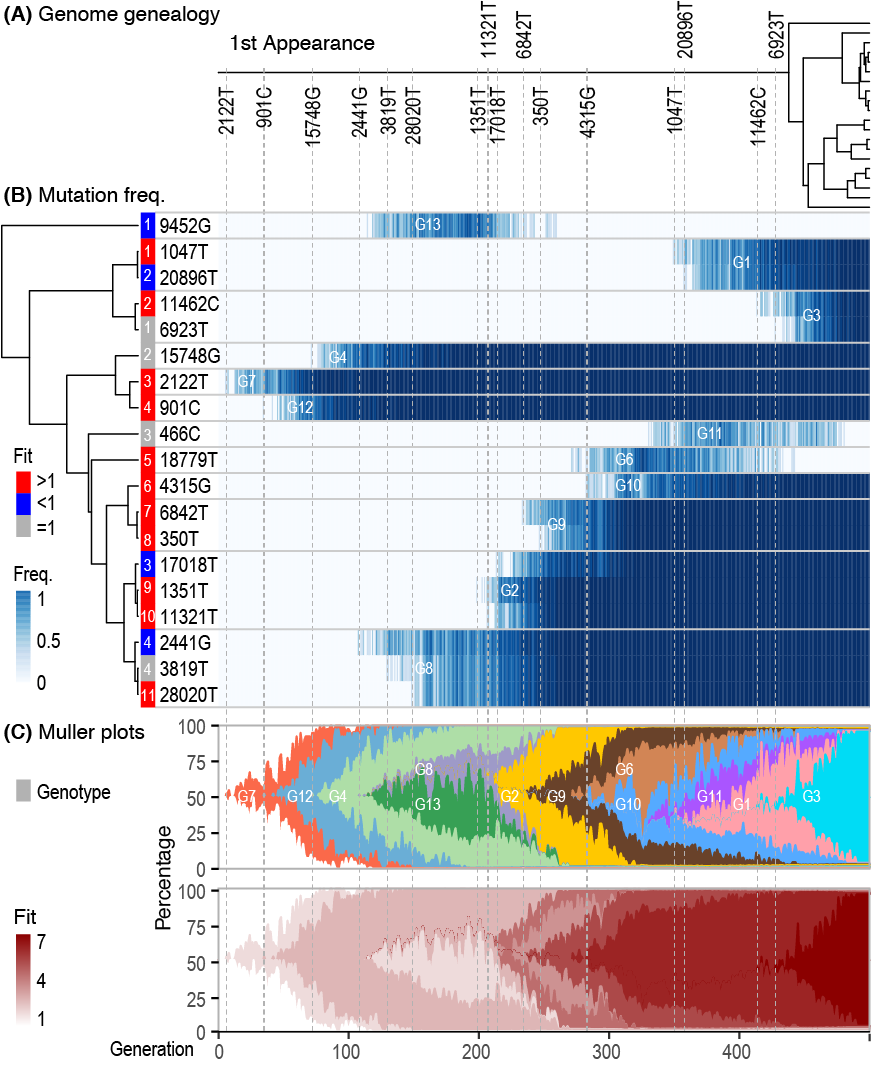
File:Cov-fig3-trace.png | Akther, Bezrucenlovas, Li, Sulkow, Di, Pante, Martin, Luft, Qiu (Sep, 2021). "Following the Trail of One Million Genomes: Footprints of SARS-CoV-2 Adaptation to Humans". '''''[https://www.biorxiv.org/content/biorxiv/early/2021/05/10/2021.05.07.443114.full.pdf BioRxiv link]''''': . Github: https://github.com/weigangq/cov-db | |||
</gallery> | |||
==Lab Resources== | ==Lab Resources & Protocols== | ||
* Github repositories: https://github.com/weigangq/?tab=repositories | * Github repositories: https://github.com/weigangq/?tab=repositories | ||
* [[Mini-Tutorals|Mini-Protocols]] (frequently used computer codes and pipelines) | * [[Mini-Tutorals|Mini-Protocols]] (frequently used computer codes and pipelines) | ||
Revision as of 21:05, 27 March 2023
Weigang Qiu, Ph.D., Professor Department of Biological Sciences Hunter College of City University of New York Belfer Research Building, Room 402 413 East 69th Street, New York, NY 10021 Office: 1-212-896-0445 Email: wqiu-at-(hunter.cuny.edu) |
Lab publications
Lyme Genomics
Evolution & Learning Algorithms
Informatics Tool Development
last update: March 20, 2023
Lab members and trainees
| Year/Period | Doctoral members & trainees | Other members & trainees |
|---|---|---|
| Current Academic Year
(Fall 2022, Spring 2023 & Summer 2023) |
|
|
| Alumni (Since Fall 2002) |
|
(published coauthors)
|
Web Apps
Apps with Collaborators
Data & GSEA associated with Polotskaia_etal_2014 (with Bargonetti Lab @Hunter)
Qiu Lab Apps
Teaching & Curricular Development
- The Quantitative Biology (QuBi) Program at Hunter
- Hunter Biology Major I (including the Bioinformatics Conentration)
- Hunter Chemistry Major 2 (including the Bioinformatics Option)
- Hunter Computer Science (including Bioinformatics Concentration)
- Hunter Mathematics (including the Quantitative Biology Concentration)
- Hunter Statistics (including the Quantitative Biology Concentration)
- QuBi module: BIOL20300 Molecular Genetics, Lab 4
- QuBi module: BIOL20300 Molecular Genetics, Lab 7
- QuBi module: BIOL30300 Cell Biology, Bioinformatics Lab (transcriptome analysis)
- Big Data (2020)
- Southwest University R course (2019)
- BIOL425 Computational Molecular Biology (2020)
- Analysis of Biological Data (2017)
- BIOL37500, Molecular Evolution (2019)
- Bioinformatics Workshop (2014)
- BIOL47120 Biomedical Genomics II (2020)
SARS-CoV-2 genome evolution
Akther, Bezrucenkovas, Sulkow, Panlasigui, Qiu, Di (April, 2020). "CoV Genome Tracker: tracing genomic footprints of Covid-19 pandemic". BioRxiv; Web app: SARS-CoV-2 Genome Tracker; Github: https://github.com/weigangq/cov-browser
Akther, Bezrucenlovas, Li, Sulkow, Di, Pante, Martin, Luft, Qiu (Sep, 2021). "Following the Trail of One Million Genomes: Footprints of SARS-CoV-2 Adaptation to Humans". BioRxiv link: . Github: https://github.com/weigangq/cov-db
Lab Resources & Protocols
- Github repositories: https://github.com/weigangq/?tab=repositories
- Mini-Protocols (frequently used computer codes and pipelines)
- Monte Carlo Club (summer projects)
- Tick handling protocols
- Hunter HPC Usage
Wiki Help
- Configuration settings list
- MediaWiki FAQ
- MediaWiki release mailing list
- Localise MediaWiki for your language
- Learn how to combat spam on your wiki
- Consult the User's Guide for information on using the wiki software.