Summer 2021: Difference between revisions
Jump to navigation
Jump to search
imported>Weigang |
imported>Weigang |
||
| Line 56: | Line 56: | ||
** Natural language models to improve structural stability (see Project 4 below) | ** Natural language models to improve structural stability (see Project 4 below) | ||
** Improve protein solubility with Solubis | ** Improve protein solubility with Solubis | ||
** Estimating Ag-Ab binding parameters with hierarchical Bayesian modeling: | |||
*** [https://www.r-bloggers.com/2016/11/hierarchical-models-with-rstan-part-1/ with R-Stan]; [https://mc-stan.org/docs/2_27/stan-users-guide/index.html Stan manual] | |||
*** [https://r-nimble.org/html_manual/cha-lightning-intro.html with R-Nimble] | |||
* Reading list | * Reading list | ||
** Greet De Baets, Joost Van Durme, Rob van der Kant, Joost Schymkowitz, Frederic Rousseau, Solubis: optimize your protein, Bioinformatics, Volume 31, Issue 15, 1 August 2015, Pages 2580–2582, https://doi.org/10.1093/bioinformatics/btv162 | ** Greet De Baets, Joost Van Durme, Rob van der Kant, Joost Schymkowitz, Frederic Rousseau, Solubis: optimize your protein, Bioinformatics, Volume 31, Issue 15, 1 August 2015, Pages 2580–2582, https://doi.org/10.1093/bioinformatics/btv162 | ||
Revision as of 16:24, 18 July 2021
Group meeting/Field trip schedule
July 2021
- Belfer housekeeping duties: During the month of July, members of Qiu lab (including the PI) will be responsible for the following:
- Performing a careful sweep of the common seating, eating, and food preparation areas each Monday, Wednesday, and Friday. The purpose of this sweep is to do some light organization/cleaning and to identify persistent cleanliness problems that should then be reported (via the lab’s PI) to myself and Elizabeth Cohn (ec2692@hunter.cuny.edu), and the collective PIs of the floor.
- Throwing away any personal food (and personal food containers) left in the refrigerators beyond 5 pm on each Friday.
- Keeping general tabs on the bathrooms of the floor to ensure cleanliness. Persistent cleanliness problems that are unreasonable for the WCMC custodial staff should be addressed by the members of the floor.
- July 2, Friday. Field trip
June 2021
- June 3, 2021 (Thursday). Summer research kickoff
- June 8, 2021 (Tuesday). 11-2
- Algorithm development (Brian)
- NLP models of protein structure (Eamen, Roman, Edgar)
- Bb transcriptomics (Niemah & Jackie)
- HIV compartmentalization (Lily)
- June 10, 2021 (Thursday). No meeting. Field day
- Location: Hackscher State Park, Long Island
- Participants: Desiree, Lily, Lia, John, Weigang
- Outcome: ~60 nymph ticks
- June 15, 2021 (Tuesday). 11-2. Lab meeting
- June 17, 2021 (Thursday). No meeting. Field day
- Location: Quabbin Reservoir, MA
- Participant: Saymon, Brian, Lia, Weigang
- Outcome: ~130 nymph ticks
- June 22, 2021 (Tuesday). 11-1
- Niemah & Jackie will work on collecting transcriptome data into Excel sheets
- Lily presented latest trees on HIV compartmentalization
- Roman will work on env gene embedding using Facebook pretrained ESM model
- June 24, 2021 (Thursday). Lab meeting 11-1
- June 30, 2021 (Wed). Lab meeting 11-1
Project 1. Borrelia genomics
- Participants: Niemah, Jackie
- Questions & Goals:
- Upgrade database, genome pipeline, and website (Lia)
- Phylogeography & evolutionary maintenance of divided genome (Saymon)
- vls evolution (with simulation) & development of immunoflorescence microsopy methods(Lily). Live imaging.
- Reading list
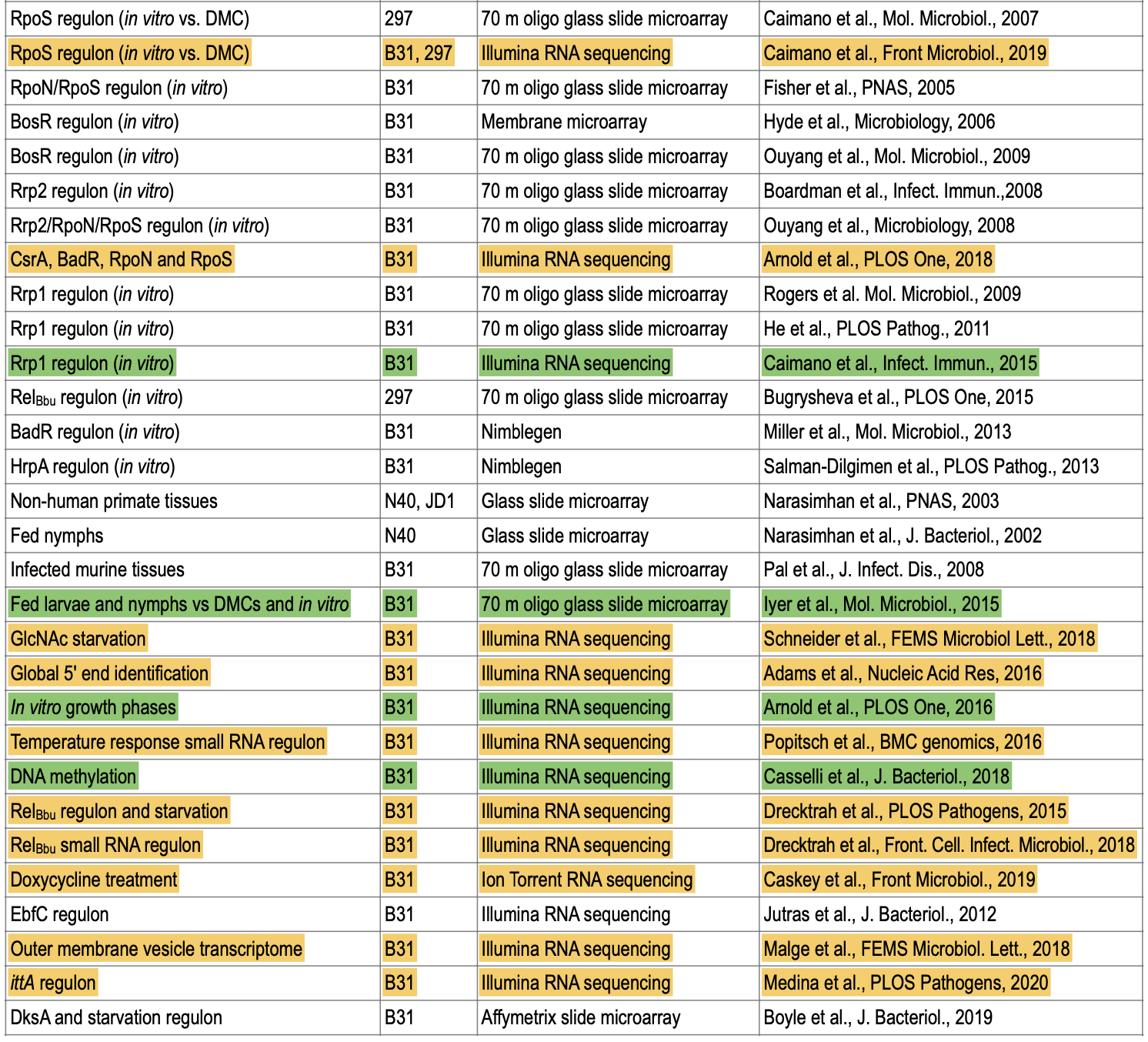
- Latest review book Lyme Disease and Relapsing Fever Spirochetes: Genomics, Molecular Biology, Host Interactions and Disease Pathogenesis. The chapter on gene regulation and transcriptomics (notice Fig 1, Fig 2, and Table 1)
- Schward et al (2021). Multipartite Genome of Lyme Disease Borrelia: Structure, Variation and Prophages
- Stevenson & Seshu (2018). Regulation of Gene and Protein Expression in the Lyme Disease Spirochete
- The bowtie model of gene regulation: Yan et al (2017). PLoS Comp Biol.
Project 2. Design algorithms for vaccines
- Participants: Dr Saad Mneimneih (CS Department), Brian
- Questions & Goals:
- Generalized algorithms for antigen with arbitrary tree shape
- Data set 1. Neutral evolution (with exponentially distributed branch lengths). Binary strings (L=100 bits) evolved from a coalescent tree of 20 leaves. Simulated with
rcoal(20); rTraitDisc; simSeq(). code from previous work - Data set 2. Two major clades. HA sequences from fluB
- Data set 3. Four major clades. Dengue
- Data set 4. Star-shaped tree, driven by recombination. OspC
- Data set 5. Multiple major clades. vls cassette in Lyme species
- Data set 1. Neutral evolution (with exponentially distributed branch lengths). Binary strings (L=100 bits) evolved from a coalescent tree of 20 leaves. Simulated with
- Combination algorithms
- Naive Bayes models to integrate immunogenicity data
- Natural language models to improve structural stability (see Project 4 below)
- Improve protein solubility with Solubis
- Estimating Ag-Ab binding parameters with hierarchical Bayesian modeling:
- Generalized algorithms for antigen with arbitrary tree shape
- Reading list
- Greet De Baets, Joost Van Durme, Rob van der Kant, Joost Schymkowitz, Frederic Rousseau, Solubis: optimize your protein, Bioinformatics, Volume 31, Issue 15, 1 August 2015, Pages 2580–2582, https://doi.org/10.1093/bioinformatics/btv162
- Degoot et al (2019). Predicting Antigenicity of Influenza A Viruses Using biophysical ideas
- Di et al (2021). Maximum antigen divergence in Lyme bacterial population
- Golovanov et al (2004). A Simple Method for Improving Protein Solubility and Long-Term Stability. J. Am. Che. Soc.
Project 3. HIV compartmentalized evolution
- Participants: Lily
- Questions and goals
- Do HIV evolve cell type tropisms within the host? Specifically, the Neural(N)-tropism vs T-cell(T)-tropism?
- Build a classifier of N-tropism HIV subtypes
- A presentation for an HIV conference in October
- Reading list
- HIV compartmentalized evolution: Evering et al (2014)
- Data sets
- ~500 sequences of env genes from 15 patients
- 2nd time point single-cell genome sequences for some of the patients
- Experimentally verified N-tropism subtypes
- Approach
- Evolutionary mechanisms: mutation, recombination, and adaptive selection
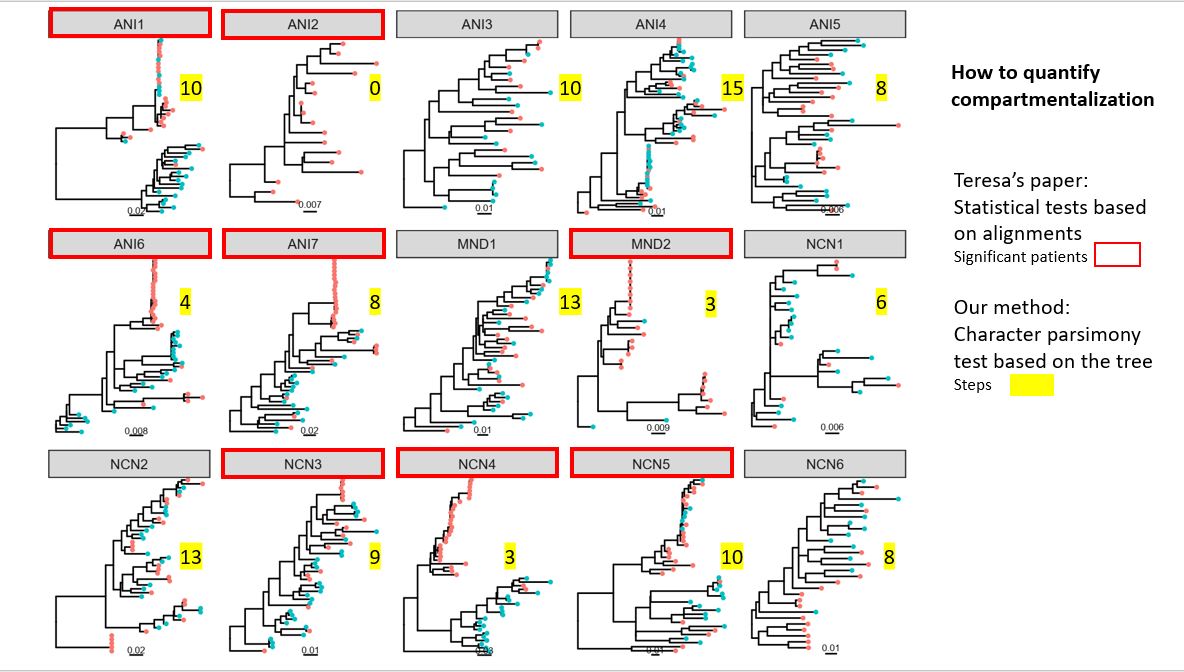
- Homoplasy index as a measure of compartmentalization? Randomization to obtain p-values of HI.
- Evolutionary rates & signature (BEAST)
- Tests of natural selection (PAML site models, branch-site models & MK analysis)
- Phylogenetic analysis: tree per individual; supertree; haplotype networks (per individual)
- Simulated compartmentalization
Project 4. Natural Language models of proteins
- Participants: Eamen, Roman, and Edgar
- Questions & Goals
- Learn, implement, and compare the existing tools
- Fine-tuning for OspC, to be integrated with the centroid algorithm
- 2nd-generation centroid design: k-means algorithm (with applications to vls, Dengue, flu B)
- Tools & Reading list
- Facebook ESM: pre-trained language models (for feature extraction)
- Step 1. implementation with colab
- Step 2. Fine-tuning with OspC seqs; extract embedding
- Step 3. Applications: classify (OspC vs VlsE), contact map (native vs synthetics), solubility
- Strodthoff et al (2020). Bioinformatics. UDSMProt: universal deep sequence models for protein classification. Source code on Github
- Rives et al (2021). PNAS. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Github repository
- Transformer: Vaswani et al (2017). Attention is All You Need Github repository for Huggingface/Transformer
- Facebook ESM: pre-trained language models (for feature extraction)